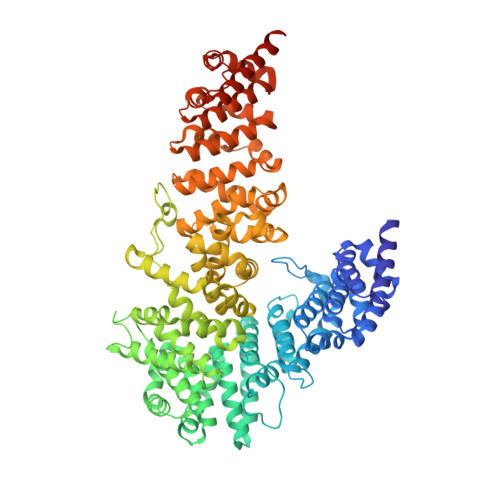
X-ray Crystal Structure of the UCS Domain-Containing UNC-45 Myosin Chaperone from Drosophila melanogaster.
Lee, C.F., Hauenstein, A.V., Fleming, J.K., Gasper, W.C., Engelke, V., Sankaran, B., Bernstein, S.I., Huxford, T.(2011) Structure 19: 397-408
- PubMed: 21397190
- DOI: https://doi.org/10.1016/j.str.2011.01.002
- Primary Citation Related Structures:
3NOW - PubMed Abstract:
UCS proteins, such as UNC-45, influence muscle contraction and other myosin-dependent motile processes. We report the first X-ray crystal structure of a UCS domain-containing protein, the UNC-45 myosin chaperone from Drosophila melanogaster (DmUNC-45). The structure reveals that the central and UCS domains form a contiguous arrangement of 17 consecutive helical layers that arrange themselves into five discrete armadillo repeat subdomains. Small-angle X-ray scattering data suggest that free DmUNC-45 adopts an elongated conformation and exhibits flexibility in solution. Protease sensitivity maps to a conserved loop that contacts the most carboxy-terminal UNC-45 armadillo repeat subdomain. Amino acid conservation across diverse UCS proteins maps to one face of this carboxy-terminal subdomain, and the majority of mutations that affect myosin-dependent cellular activities lie within or around this region. Our crystallographic, biophysical, and biochemical analyses suggest that DmUNC-45 function is afforded by its flexibility and by structural integrity of its UCS domain.
- Department of Biology, San Diego State University, 5500 Campanile Drive, San Diego, CA 92182-1030, USA.
Organizational Affiliation: