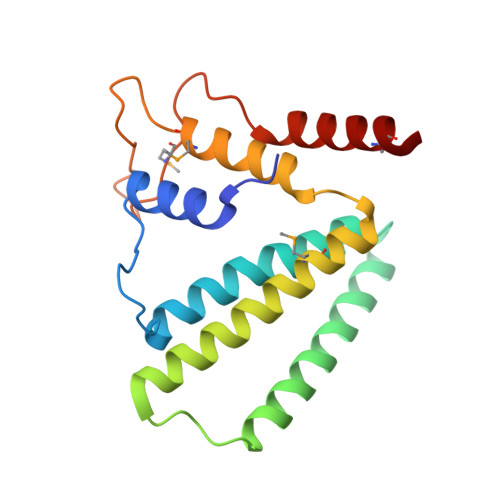
Structure of a putative NTP pyrophosphohydrolase: YP_001813558.1 from Exiguobacterium sibiricum 255-15.
Han, G.W., Elsliger, M.A., Yeates, T.O., Xu, Q., Murzin, A.G., Krishna, S.S., Jaroszewski, L., Abdubek, P., Astakhova, T., Axelrod, H.L., Carlton, D., Chen, C., Chiu, H.J., Clayton, T., Das, D., Deller, M.C., Duan, L., Ernst, D., Feuerhelm, J., Grant, J.C., Grzechnik, A., Jin, K.K., Johnson, H.A., Klock, H.E., Knuth, M.W., Kozbial, P., Kumar, A., Lam, W.W., Marciano, D., McMullan, D., Miller, M.D., Morse, A.T., Nigoghossian, E., Okach, L., Reyes, R., Rife, C.L., Sefcovic, N., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Hodgson, K.O., Wooley, J., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1237-1244
- PubMed: 20944217 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309110025534
- Primary Citation Related Structures:
3NL9 - PubMed Abstract:
The crystal structure of a putative NTPase, YP_001813558.1 from Exiguobacterium sibiricum 255-15 (PF09934, DUF2166) was determined to 1.78 Å resolution. YP_001813558.1 and its homologs (dimeric dUTPases, MazG proteins and HisE-encoded phosphoribosyl ATP pyrophosphohydrolases) form a superfamily of all-α-helical NTP pyrophosphatases. In dimeric dUTPase-like proteins, a central four-helix bundle forms the active site. However, in YP_001813558.1, an unexpected intertwined swapping of two of the helices that compose the conserved helix bundle results in a `linked dimer' that has not previously been observed for this family. Interestingly, despite this novel mode of dimerization, the metal-binding site for divalent cations, such as magnesium, that are essential for NTPase activity is still conserved. Furthermore, the active-site residues that are involved in sugar binding of the NTPs are also conserved when compared with other α-helical NTPases, but those that recognize the nucleotide bases are not conserved, suggesting a different substrate specificity.
- Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.
Organizational Affiliation: