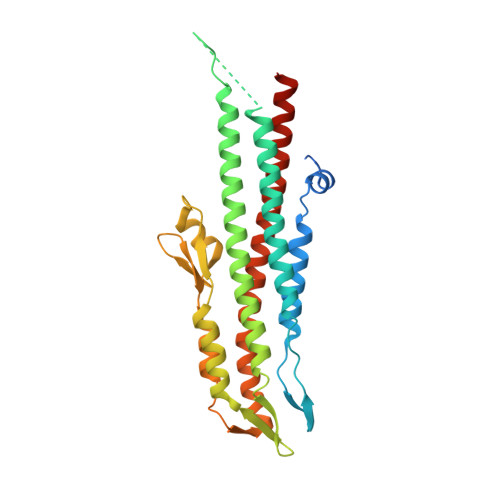
Near-atomic resolution analysis of BipD, a component of the type III secretion system of Burkholderia pseudomallei.
Pal, M., Erskine, P.T., Gill, R.S., Wood, S.P., Cooper, J.B.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 990-993
- PubMed: 20823511 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309110026333
- Primary Citation Related Structures:
3NFT - PubMed Abstract:
Burkholderia pseudomallei, the causative agent of melioidosis, possesses a type III protein secretion apparatus that is similar to those found in Salmonella and Shigella. A major function of these secretion systems is to inject virulence-associated proteins into target cells of the host organism. The bipD gene of B. pseudomallei encodes a secreted virulence factor that is similar in sequence and is most likely to be functionally analogous to IpaD from Shigella and SipD from Salmonella. Proteins in this family are thought to act as extracellular chaperones at the tip of the secretion needle to help the hydrophobic translocator proteins enter the target cell membrane, where they form a pore and may also link the translocon pore with the secretion needle. BipD has been crystallized in a monoclinic crystal form that diffracted X-rays to 1.5 A resolution and the structure was refined to an R factor of 16.1% and an Rfree of 19.8% at this resolution. The putative dimer interface that was observed in previous crystal structures was retained and a larger surface area was buried in the new crystal form.
- School of Biological Sciences, University of Southampton, Bassett Crescent East, Southampton SO16 7PX, England.
Organizational Affiliation: