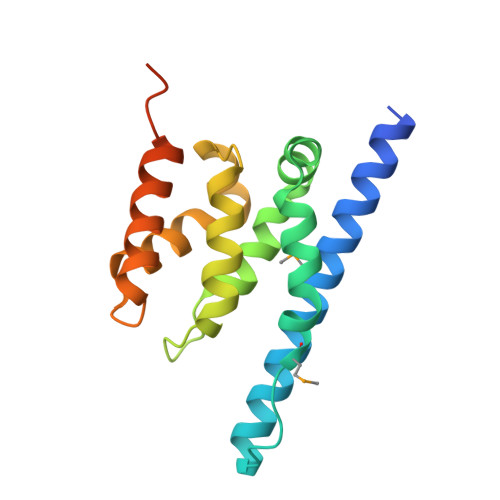
The Transcription Factor Spn1 Regulates Gene Expression via a Highly Conserved Novel Structural Motif.
Pujari, V., Radebaugh, C.A., Chodaparambil, J.V., Muthurajan, U.M., Almeida, A.R., Fischbeck, J.A., Luger, K., Stargell, L.A.(2010) J Mol Biology 404: 1-15
- PubMed: 20875428 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.09.040
- Primary Citation Related Structures:
3NFQ - PubMed Abstract:
Spn1/Iws1 plays essential roles in the regulation of gene expression by RNA polymerase II (RNAPII), and it is highly conserved in organisms ranging from yeast to humans. Spn1 physically and/or genetically interacts with RNAPII, TBP (TATA-binding protein), TFIIS (transcription factor IIS), and a number of chromatin remodeling factors (Swi/Snf and Spt6). The central domain of Spn1 (residues 141-305 out of 410) is necessary and sufficient for performing the essential functions of SPN1 in yeast cells. Here, we report the high-resolution (1.85 Å) crystal structure of the conserved central domain of Saccharomyces cerevisiae Spn1. The central domain is composed of eight α-helices in a right-handed superhelical arrangement and exhibits structural similarity to domain I of TFIIS. A unique structural feature of Spn1 is a highly conserved loop, which defines one side of a pronounced cavity. The loop and the other residues forming the cavity are highly conserved at the amino acid level among all Spn1 family members, suggesting that this is a signature motif for Spn1 orthologs. The locations and the molecular characterization of temperature-sensitive mutations in Spn1 indicate that the cavity is a key attribute of Spn1 that is critical for its regulatory functions during RNAPII-mediated transcriptional activity.
- Department of Biochemistry and Molecular Biology, Colorado State University, Fort Collins, CO 80523-1870, USA.
Organizational Affiliation: