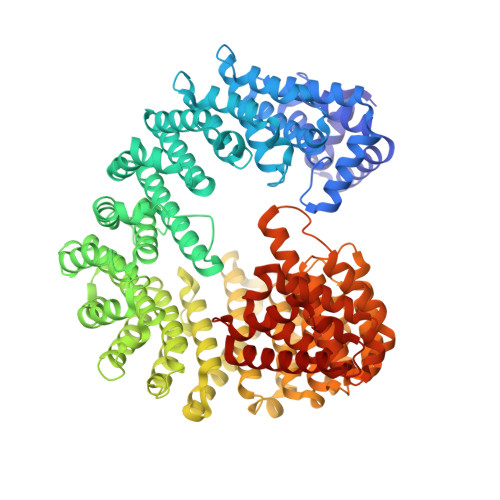
Quantitative Structural Analysis of Importin-beta Flexibility: Paradigm for Solenoid Protein Structures
Forwood, J.K., Lange, A., Zachariae, U., Marfori, M., Preast, C., Grubmuller, H., Stewart, M., Corbett, A.H., Kobe, B.(2010) Structure 18: 1171-1183
- PubMed: 20826343 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2010.06.015
- Primary Citation Related Structures:
3ND2 - PubMed Abstract:
The structure of solenoid proteins facilitates a higher degree of flexibility than most folded proteins. In importin-β, a nuclear import factor built from 19 tandem HEAT repeats, flexibility plays a crucial role in allowing interactions with a range of different partners. We present a comprehensive analysis of importin-β flexibility based on a number of different approaches. We determined the crystal structure of unliganded Saccharomyces cerevisiae importin-β (Kap95) to allow a quantitative comparison with importin-β bound to different partners. Complementary mutagenesis, small angle X-ray scattering and molecular dynamics studies suggest that the protein samples several conformations in solution. The analyses suggest the flexibility of the solenoid is generated by cumulative small movements along its length. Importin-β illustrates how solenoid proteins can orchestrate protein interactions in many cellular pathways.
- School of Biomedical Sciences, Charles Sturt University, Wagga Wagga, New South Wales 2650, Australia. jforwood@csu.edu.au
Organizational Affiliation: