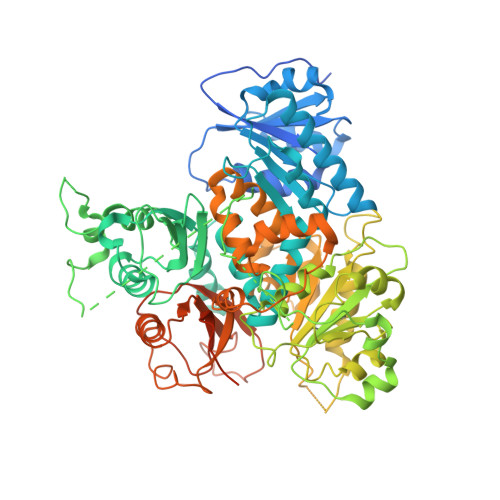
Structure of the gating ring from the human large-conductance Ca(2+)-gated K(+) channel.
Wu, Y., Yang, Y., Ye, S., Jiang, Y.(2010) Nature 466: 393-397
- PubMed: 20574420 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature09252
- Primary Citation Related Structures:
3NAF - PubMed Abstract:
Large-conductance Ca(2+)-gated K(+) (BK) channels are essential for many biological processes such as smooth muscle contraction and neurotransmitter release. This group of channels can be activated synergistically by both voltage and intracellular Ca(2+), with the large carboxy-terminal intracellular portion being responsible for Ca(2+) sensing. Here we present the crystal structure of the entire cytoplasmic region of the human BK channel in a Ca(2+)-free state. The structure reveals four intracellular subunits, each comprising two tandem RCK domains, assembled into a gating ring similar to that seen in the MthK channel and probably representing its physiological assembly. Three Ca(2+) binding sites including the Ca(2+) bowl are mapped onto the structure based on mutagenesis data. The Ca(2+) bowl, located within the second RCK domain, forms an EF-hand-like motif and is strategically positioned close to the assembly interface between two subunits. The other two Ca(2+) (or Mg(2+)) binding sites, Asp 367 and Glu 374/Glu 399, are located on the first RCK domain. The Asp 367 site has high Ca(2+) sensitivity and is positioned in the groove between the amino- and carboxy-terminal subdomains of RCK1, whereas the low-affinity Mg(2+)-binding Glu 374/Glu 399 site is positioned on the upper plateau of the gating ring and close to the membrane. Our structure also contains the linker connecting the transmembrane and intracellular domains, allowing us to dock a voltage-gated K(+) channel pore of known structure onto the gating ring with reasonable accuracy and generate a structural model for the full BK channel.
- Department of Physiology, University of Texas Southwestern Medical Center, Dallas, Texas 75390-9040, USA.
Organizational Affiliation: