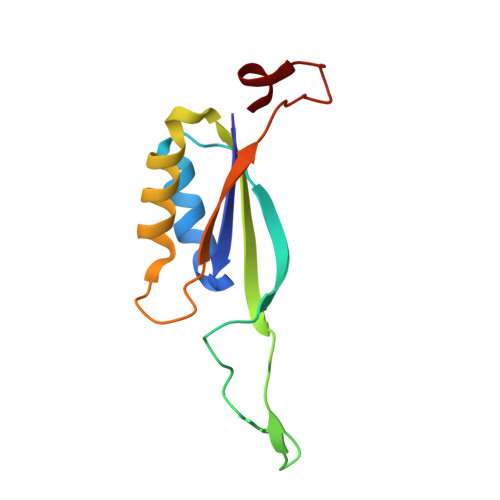
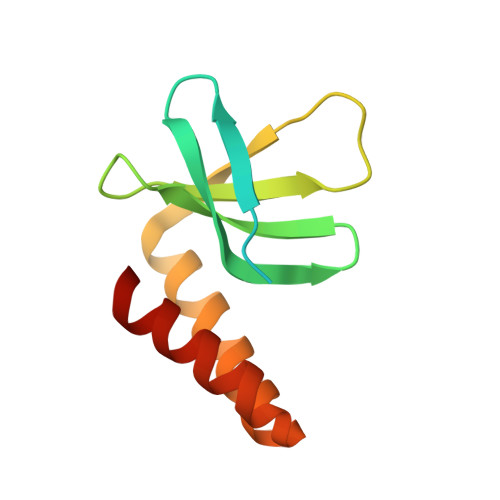
Crystal structure of the cyanobacterial signal transduction protein PII in complex with PipX.
Zhao, M.-X., Jiang, Y.-L., Xu, B.-Y., Chen, Y.-X., Zhang, C.-C., Zhou, C.-Z.(2010) J Mol Biology 402: 552-559
- PubMed: 20708625 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.08.006
- Primary Citation Related Structures:
3N5B - PubMed Abstract:
P(II) proteins are highly conserved signal transducers in bacteria, archaea, and plants. They have a large flexible loop (T-loop) that adopts different conformations after covalent modification or binding to different effectors to regulate the functions of diverse protein partners. The P(II) partner PipX (P(II)interaction protein X), first identified from Synechococcus sp. PCC 7942, exists uniquely in cyanobacteria. PipX also interacts with the cyanobacterial global nitrogen regulator NtcA. The mutually exclusive binding of P(II) and NtcA by PipX in a 2-oxoglutarate (2-OG)-dependent manner enables P(II) to indirectly regulate the transcriptional activity of NtcA. However, the structural basis for these exclusive interactions remains unknown. We solved the crystal structure of the P(II)-PipX complex from the filamentous cyanobacterium Anabaena sp. PCC 7120 at 1.90 Å resolution. A homotrimeric P(II) captures three subunits of PipX through the T-loops. Similar to P(II) from Synechococcus, the core structure consists of an antiparallel β-sheet with four β-strands and two α-helices at the lateral surface. PipX adopts a novel structure composed of five twisted antiparallel β-strands and two α-helices, which is reminiscent of the P(II) structure. The T-loop of each P(II) subunit extends from the core structure as an antenna that is stabilized at the cleft between two PipX monomers via hydrogen bonds. In addition, the interfaces between the β-sheets of PipX and P(II) core structures partially contribute to complex formation. Comparative structural analysis indicated that PipX and 2-OG share a common binding site that overlaps with the 14 signature residues of cyanobacterial P(II) proteins. Our structure of PipX and the recently solved NtcA structure enabled us to propose a putative model for the NtcA-PipX complex. Taken together, these findings provide structural insights into how P(II) regulates the transcriptional activity of NtcA via PipX upon accumulation of the metabolite 2-OG.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, People's Republic of China.
Organizational Affiliation: