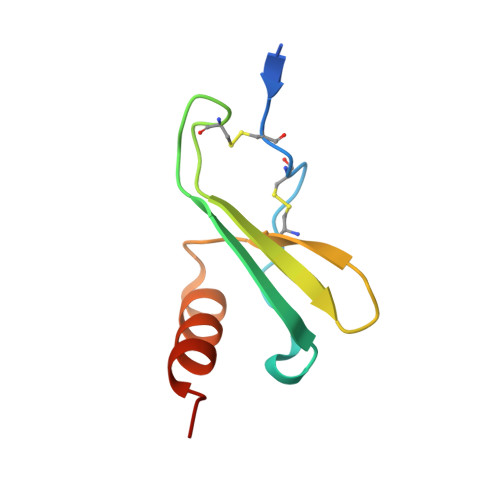
A Model of GAG/MIP-2/CXCR2 Interfaces and Its Functional Effects.
Rajasekaran, D., Keeler, C., Syed, M.A., Jones, M.C., Harrison, J.K., Wu, D., Bhandari, V., Hodsdon, M.E., Lolis, E.J.(2012) Biochemistry 51: 5642-5654
- PubMed: 22686371 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi3001566
- Primary Citation Related Structures:
3N52 - PubMed Abstract:
MIP-2/CXCL2 is a murine chemokine related to human chemokines that possesses the Glu-Leu-Arg (ELR) activation motif and activates CXCR2 for neutrophil chemotaxis. We determined the structure of MIP-2 to 1.9 Å resolution and created a model with its murine receptor CXCR2 based on the coordinates of human CXCR4. Chemokine-induced migration of cells through specific G-protein coupled receptors is regulated by glycosaminoglycans (GAGs) that oligomerize chemokines. MIP-2 GAG-binding residues were identified that interact with heparin disaccharide I-S by NMR spectroscopy. A model GAG/MIP-2/CXCR2 complex that supports a 2:2 complex between chemokine and receptor was created. Mutants of these disaccharide-binding residues were made and tested for heparin binding, in vitro neutrophil chemotaxis, and in vivo neutrophil recruitment to the mouse peritoneum and lung. The mutants have a 10-fold decrease in neutrophil chemotaxis in vitro. There is no difference in neutrophil recruitment between wild-type MIP-2 and mutants in the peritoneum, but all activity of the mutants is lost in the lung, supporting the concept that GAG regulation of chemokines is tissue-dependent.
- Departments of Pharmacology, Yale University School of Medicine, New Haven, CT 06520-8066, USA.
Organizational Affiliation: