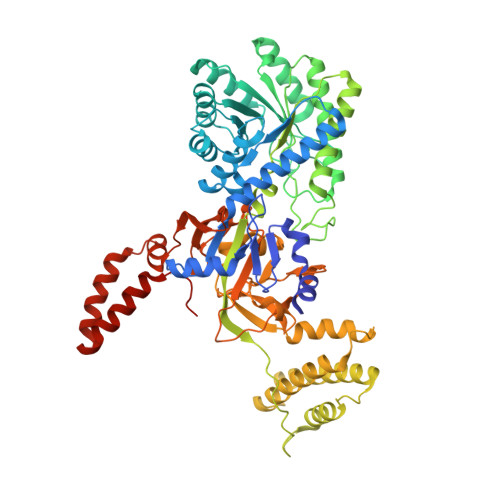
Evolution of substrate specificity within a diverse family of beta/alpha-barrel-fold basic amino acid decarboxylases: X-ray structure determination of enzymes with specificity for L-arginine and carboxynorspermidine.
Deng, X., Lee, J., Michael, A.J., Tomchick, D.R., Goldsmith, E.J., Phillips, M.A.(2010) J Biological Chem 285: 25708-25719
- PubMed: 20534592 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.121137
- Primary Citation Related Structures:
3N29, 3N2O - PubMed Abstract:
Pyridoxal 5'-phosphate (PLP)-dependent basic amino acid decarboxylases from the beta/alpha-barrel-fold class (group IV) exist in most organisms and catalyze the decarboxylation of diverse substrates, essential for polyamine and lysine biosynthesis. Herein we describe the first x-ray structure determination of bacterial biosynthetic arginine decarboxylase (ADC) and carboxynorspermidine decarboxylase (CANSDC) to 2.3- and 2.0-A resolution, solved as product complexes with agmatine and norspermidine. Despite low overall sequence identity, the monomeric and dimeric structures are similar to other enzymes in the family, with the active sites formed between the beta/alpha-barrel domain of one subunit and the beta-barrel of the other. ADC contains both a unique interdomain insertion (4-helical bundle) and a C-terminal extension (3-helical bundle) and it packs as a tetramer in the asymmetric unit with the insertions forming part of the dimer and tetramer interfaces. Analytical ultracentrifugation studies confirmed that the ADC solution structure is a tetramer. Specificity for different basic amino acids appears to arise primarily from changes in the position of, and amino acid replacements in, a helix in the beta-barrel domain we refer to as the "specificity helix." Additionally, in CANSDC a key acidic residue that interacts with the distal amino group of other substrates is replaced by Leu(314), which interacts with the aliphatic portion of norspermidine. Neither product, agmatine in ADC nor norspermidine in CANSDC, form a Schiff base to pyridoxal 5'-phosphate, suggesting that the product complexes may promote product release by slowing the back reaction. These studies provide insight into the structural basis for the evolution of novel function within a common structural-fold.
- Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, Texas 75390-9041, USA.
Organizational Affiliation: