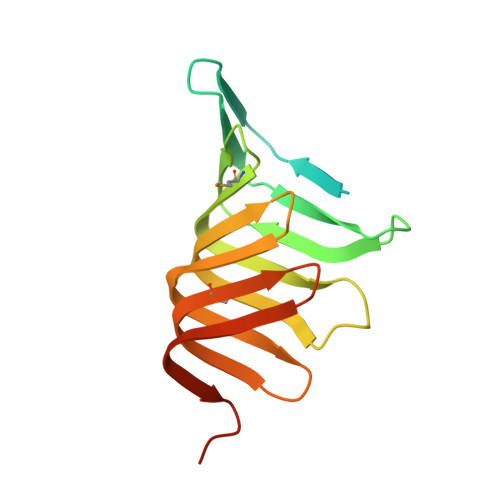
Structure and functional analysis of LptC, a conserved membrane protein involved in the lipopolysaccharide export pathway in Escherichia coli.
Tran, A.X., Dong, C., Whitfield, C.(2010) J Biological Chem 285: 33529-33539
- PubMed: 20720015 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.144709
- Primary Citation Related Structures:
3MY2 - PubMed Abstract:
LptC is a conserved bitopic inner membrane protein from Escherichia coli involved in the export of lipopolysaccharide from its site of synthesis in the cytoplasmic membrane to the outer membrane. LptC forms a complex with the ATP-binding cassette transporter, LptBFG, which is thought to facilitate the extraction of lipopolysaccharide from the inner membrane and release it into a translocation pathway that includes the putative periplasmic chaperone LptA. Cysteine modification experiments established that the catalytic domain of LptC is oriented toward the periplasm. The structure of the periplasmic domain is described at a resolution of 2.2-Å from x-ray crystallographic data. The periplasmic domain of LptC consists of a twisted boat structure with two β-sheets in apposition to each other. The β-sheets contain seven and eight antiparallel β-strands, respectively. This structure bears a high degree of resemblance to the crystal structure of LptA. Like LptA, LptC binds lipopolysaccharide in vitro. In vitro, LptA can displace lipopolysaccharide from LptC (but not vice versa), consistent with their locations and their proposed placement in a unidirectional export pathway.
- From the Department of Molecular and Cellular Biology, University of Guelph, Guelph, Ontario N1G 2W1, Canada.
Organizational Affiliation: