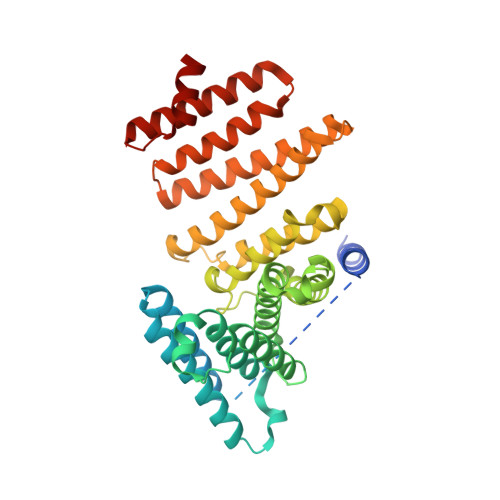
The hypoxic regulator of sterol synthesis nro1 is a nuclear import adaptor.
Yeh, T.L., Lee, C.Y., Amzel, L.M., Espenshade, P.J., Bianchet, M.A.(2011) Structure 19: 503-514
- PubMed: 21481773
- DOI: https://doi.org/10.1016/j.str.2011.01.017
- Primary Citation Related Structures:
3MSV, 6E0T - PubMed Abstract:
Fission yeast protein Sre1, the homolog of the mammalian sterol regulatory element-binding protein (SREBP), is a hypoxic transcription factor required for sterol homeostasis and low-oxygen growth. Nro1 regulates the stability of the N-terminal transcription factor domain of Sre1 (Sre1N) by inhibiting the action of the prolyl 4-hydroxylase-like Ofd1 in an oxygen-dependent manner. The crystal structure of Nro1 determined at 2.2 Å resolution shows an all-α-helical fold that can be divided into two domains: a small N-terminal domain, and a larger C-terminal HEAT-repeat domain. Follow-up studies showed that Nro1 defines a new class of nuclear import adaptor that functions both in Ofd1 nuclear localization and in the oxygen-dependent inhibition of Ofd1 to control the hypoxic response.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, 725 N Wolfe Street, Baltimore, MD 21205, USA.
Organizational Affiliation: