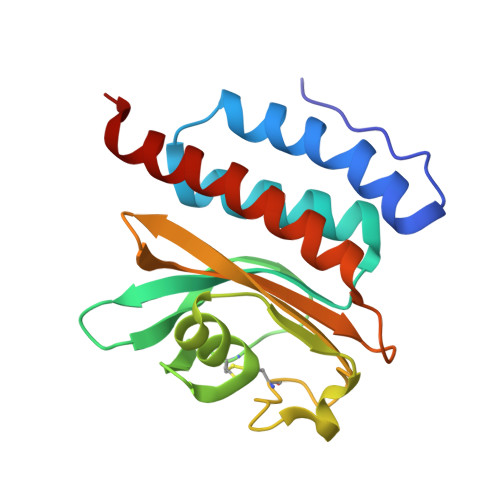
Structural and Biochemical Characterization of Free Methionine-R-sulfoxide Reductase from Neisseria meningitidis.
Gruez, A., Libiad, M., Boschi-Muller, S., Branlant, G.(2010) J Biological Chem 285: 25033-25043
- PubMed: 20489204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.134528
- Primary Citation Related Structures:
3MMH - PubMed Abstract:
A new family of methionine-sulfoxide reductase (Msr) was recently described. The enzyme, named fRMsr, selectively reduces the R isomer at the sulfoxide function of free methionine sulfoxide (Met-R-O). The fRMsrs belong to the GAF fold family. They represent the first GAF domain to show enzymatic activity. Two other Msr families, MsrA and MsrB, were already known. MsrA and MsrB reduce free Met-S-O and Met-R-O, respectively, but exhibit higher catalytic efficiency toward Met-O within a peptide or a protein context. The fold of the three families differs. In the present work, the crystal structure of the fRMsr from Neisseria meningitidis has been determined in complex with S-Met-R-O. Based on biochemical and kinetic data as well as genomic analyses, Cys(118) is demonstrated to be the catalytic Cys on which a sulfenic acid is formed. All of the structural factors involved in the stereoselectivity of the l-Met-R-O binding were identified and account for why Met-S-O, DMSO, and a Met-O within a peptide are not substrates. Taking into account the structural, enzymatic, and biochemical information, a scenario of the catalysis for the reductase step is proposed. Based on the thiol content before and after Met-O reduction and the stoichiometry of Met formed per subunit of wild type and Cys-to-Ala mutants, a scenario of the recycling process of the N. meningitidis fRMsr is proposed. All of the biochemical, enzymatic, and structural properties of the N. meningitidis fRMsr are compared with those of MsrA and MsrB and are discussed in terms of the evolution of function of the GAF domain.
- Faculté des Sciences et Technologies, AREMS, UMR CNRS-UHP 7214, Nancy Université, Bld des Aiguillettes, BP 70239, 54506 Vandoeuvre-les-Nancy, France.
Organizational Affiliation: