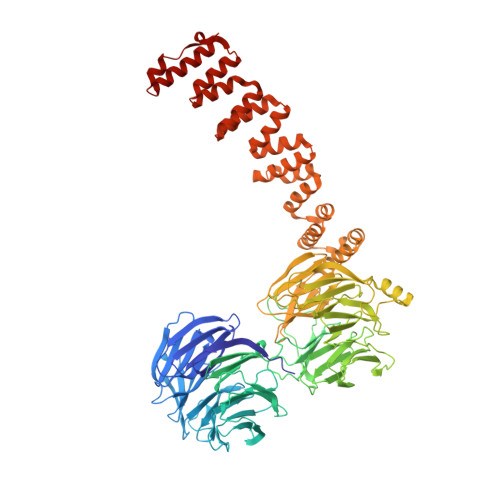
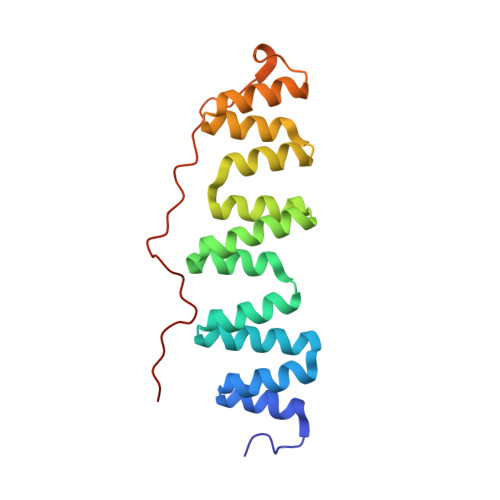
Structure of coatomer cage proteins and the relationship among COPI, COPII, and clathrin vesicle coats.
Lee, C., Goldberg, J.(2010) Cell 142: 123-132
- PubMed: 20579721 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2010.05.030
- Primary Citation Related Structures:
3MKQ, 3MKR - PubMed Abstract:
COPI-coated vesicles form at the Golgi apparatus from two cytosolic components, ARF G protein and coatomer, a heptameric complex that can polymerize into a cage to deform the membrane into a bud. Although coatomer shares a common evolutionary origin with COPII and clathrin vesicle coat proteins, the architectural relationship among the three cages is unclear. Strikingly, the alphabeta'-COP core of coatomer crystallizes as a triskelion in which three copies of a beta'-COP beta-propeller domain converge through their axial ends. We infer that the trimer constitutes the vertex of the COPI cage. Our model proposes that the COPI cage is intermediate in design between COPII and clathrin: COPI shares with clathrin an arrangement of three curved alpha-solenoid legs radiating from a common center, and COPI shares with COPII highly similar vertex interactions involving the axial ends of beta-propeller domains.
- Howard Hughes Medical Institute, Memorial Sloan-Kettering Cancer Center, 1275 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: