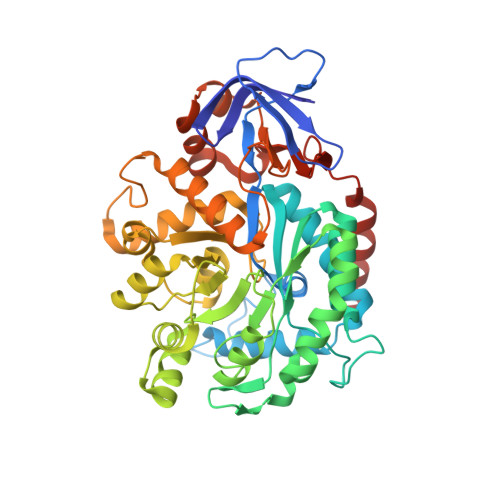
Structure of N-Formimino-l-glutamate Iminohydrolase from Pseudomonas aeruginosa.
Fedorov, A.A., Marti-Arbona, R., Nemmara, V.V., Hitchcock, D., Fedorov, E.V., Almo, S.C., Raushel, F.M.(2015) Biochemistry 54: 890-897
- PubMed: 25559274 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi501299y
- Primary Citation Related Structures:
3MDU, 3MDW, 4RZB - PubMed Abstract:
N-Formimino-l-glutamate iminohydrolase (HutF), from Pseudomonas aeruginosa with a locus tag of Pa5106 ( gi|15600299 ), is a member of the amidohydrolase superfamily. This enzyme catalyzes the deamination of N-formimino-l-glutamate to N-formyl-l-glutamate and ammonia in the histidine degradation pathway. The crystal structure of Pa5106 was determined in the presence of the inhibitors N-formimino-l-aspartate and N-guanidino-l-glutaric acid at resolutions of 1.9 and 1.4 Å, respectively. The structure of an individual subunit is composed of two domains with the larger domain folding as a distorted (β/α)8-barrel. The (β/α)8-barrel domain is composed of eight β-strands flanked by 11 α-helices, whereas the smaller domain is made up of eight β-strands. The active site of Pa5106 contains a single zinc atom that is coordinated by His-56, His-58, His-232, and Asp-320. The nucleophilic solvent water molecule coordinates with the zinc atom at a distance of 2.0 Å and is hydrogen bonded to Asp-320 and His-269. The α-carboxylate groups of both inhibitors are hydrogen bonded to the imidazole moiety of His-206, the hydroxyl group of Tyr-121, and the side chain amide group of Gln-61. The side chain carboxylate groups of the two inhibitors are ion-paired with the guanidino groups of Arg-209 and Arg-82. Computational docking of high-energy tetrahedral intermediate forms of the substrate, N-formimino-l-glutamate, to the three-dimensional structure of Pa5106 suggests that this compound likely undergoes a re-faced nucleophilic attack at the formimino group by the metal-bound hydroxide. A catalytic mechanism of the reaction catalyzed by Pa5106 is proposed.
- Department of Chemistry and ‡Department of Biochemistry & Biophysics, Texas A&M University , College Station, Texas 77843, United States.
Organizational Affiliation: