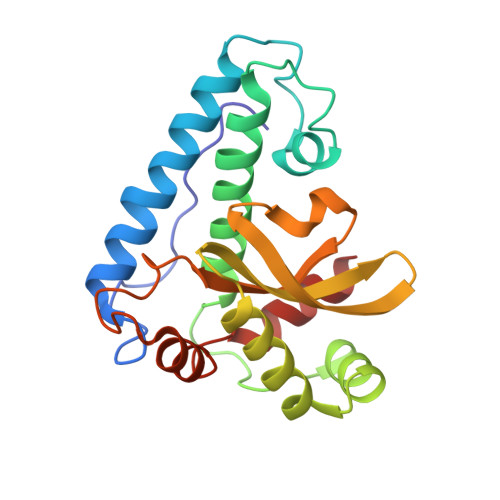
Manganese superoxide dismutase from Thermus thermophilus. A structural model refined at 1.8 A resolution.
Ludwig, M.L., Metzger, A.L., Pattridge, K.A., Stallings, W.C.(1991) J Mol Biology 219: 335-358
- PubMed: 2038060 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(91)90569-r
- Primary Citation Related Structures:
3MDS - PubMed Abstract:
The structure of Mn(III) superoxide dismutase (Mn(III)SOD) from Thermus thermophilus, a tetramer of chains 203 residues in length, has been refined by restrained least-squares methods. The R-factor [formula: see text] for the 54,056 unique reflections measured between 10.0 and 1.8 A (96% of all possible reflections) is 0.176 for a model comprising the protein dimer and 180 bound solvents, the asymmetric unit of the P4(1)2(1)2 cell. The monomer chain forms two domains as determined by distance plots: the N-terminal domain is dominated by two long antiparallel helices (residues 21 to 45 and 69 to 89) and the C-terminal domain (residues 100 to 203) is an alpha + beta structure including a three-stranded sheet. Features that may be important for the folding and function of this MnSOD include: (1) a cis-proline in a turn preceding the first long helix; (2) a residue inserted at position 30 that distorts the helix near the first Mn ligand; and (3) the locations of glycine and proline residues in the domain connector (residues 92 to 99) and in the vicinity of the short cross connection (residues 150 to 159) that links two strands of the beta-sheet. Domain-domain contacts include salt bridges between arginine residues and acidic side chains, an extensive hydrophobic interface, and at least ten hydrogen-bonded interactions. The tetramer possesses 222 symmetry but is held together by only two types of interfaces. The dimer interface at the non-crystallographic dyad is extensive (1000 A2 buried surface/monomer) and incorporates 17 trapped or structural solvents. The dimer interface at the crystallographic dyad buries fewer residues (750 A2/monomer) and resembles a snap fastener in which a type I turn thrusts into a hydrophobic basket formed by a ring of helices in the opposing chain. Each of the metal sites is fully occupied, with the Mn(III) five-co-ordinate in trigonal bipyramidal geometry. One of the axial ligands is solvent; the four protein ligands are His28, His83, Asp166 and His170. Surrounding the metal-ligand cluster is a shell of predominantly hydrophobic residues from both chains of the asymmetric unit (Phe86A, Trp87A, Trp132A, Trp168A, Tyr183A, Tyr172B, Tyr173B), and both chains collaborate in the formation of a solvent-lined channel that terminates at Tyr36 and His32 near the metal ion and is presumed to be the path by which substrate or other inner-sphere ligands reach the metal.(ABSTRACT TRUNCATED AT 400 WORDS)
- Department of Biological Chemistry, University of Michigan, Ann Arbor 48109.
Organizational Affiliation: