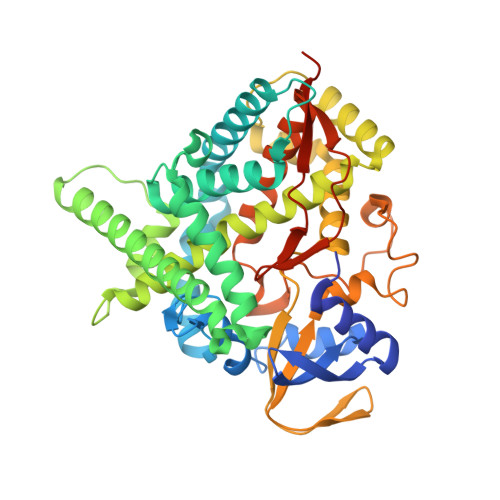
Structural basis of drug binding to CYP46A1, an enzyme that controls cholesterol turnover in the brain.
Mast, N., Charvet, C., Pikuleva, I.A., Stout, C.D.(2010) J Biological Chem 285: 31783-31795
- PubMed: 20667828 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.143313
- Primary Citation Related Structures:
3MDM, 3MDR, 3MDT, 3MDV - PubMed Abstract:
Cytochrome P450 46A1 (CYP46A1) initiates the major pathway of cholesterol elimination from the brain and thereby controls cholesterol turnover in this organ. We determined x-ray crystal structures of CYP46A1 in complex with four structurally distinct pharmaceuticals; antidepressant tranylcypromine (2.15 Å), anticonvulsant thioperamide (1.65 Å), antifungal voriconazole (2.35 Å), and antifungal clotrimazole (2.50 Å). All four drugs are nitrogen-containing compounds that have nanomolar affinity for CYP46A1 in vitro yet differ in size, shape, hydrophobicity, and type of the nitrogen ligand. Structures of the co-complexes demonstrate that each drug binds in a single orientation to the active site with tranylcypromine, thioperamide, and voriconazole coordinating the heme iron via their nitrogen atoms and clotrimazole being at a 4 Å distance from the heme iron. We show here that clotrimazole is also a substrate for CYP46A1. High affinity for CYP46A1 is determined by a set of specific interactions, some of which were further investigated by solution studies using structural analogs of the drugs and the T306A CYP46A1 mutant. Collectively, our results reveal how diverse inhibitors can be accommodated in the CYP46A1 active site and provide an explanation for the observed differences in the drug-induced spectral response. Co-complexes with tranylcypromine, thioperamide, and voriconazole represent the first structural characterization of the drug binding to a P450 enzyme.
- Department of Ophthalmology and Visual Sciences, Case Western Reserve University, Cleveland, Ohio 44106, USA.
Organizational Affiliation: