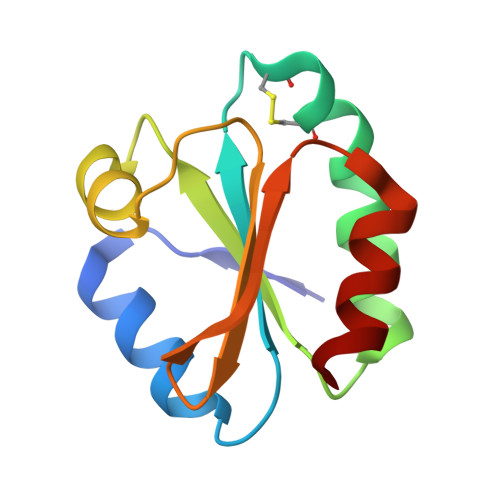
Crystal structure of human thioredoxin revealing an unraveled helix and exposed S-nitrosation site.
Weichsel, A., Kem, M., Montfort, W.R.(2010) Protein Sci 19: 1801-1806
- PubMed: 20662007 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.455
- Primary Citation Related Structures:
3M9J, 3M9K - PubMed Abstract:
Thioredoxins reduce disulfide bonds and other thiol modifications in all cells using a CXXC motif. Human thioredoxin 1 is unusual in that it codes for an additional three cysteines in its 105 amino acid sequence, each of which have been implicated in other reductive activities. Cys 62 and Cys 69 are buried in the protein interior and lie at either end of a short helix (helix 3), and yet can disulfide link under oxidizing conditions. Cys 62 is readily S-nitrosated, giving rise to a SNO modification, which is also buried. Here, we present two crystal structures of the C69S/C73S mutant protein under oxidizing (1.5 A) and reducing (1.1 A) conditions. In the oxidized structure, helix 3 is unraveled and displays a new conformation that is stabilized by a series of new hydrogen bonds and a disulfide link with Cys 62 in a neighboring molecule. The new conformation provides an explanation for how a completely buried residue can participate in SNO exchange reactions.
- Department of Chemistry and Biochemistry, University of Arizona, Tucson, Arizona 85721, USA.
Organizational Affiliation: