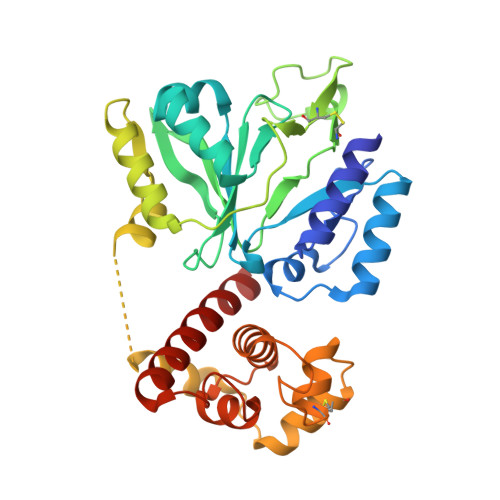
The archaeo-eukaryotic primase of plasmid pRN1 requires a helix bundle domain for faithful primer synthesis
Beck, K., Vannini, A., Cramer, P., Lipps, G.(2010) Nucleic Acids Res 38: 6707-6718
- PubMed: 20511586
- DOI: https://doi.org/10.1093/nar/gkq447
- Primary Citation Related Structures:
3M1M - PubMed Abstract:
The plasmid pRN1 encodes for a multifunctional replication protein with primase, DNA polymerase and helicase activity. The minimal region required for primase activity encompasses amino-acid residues 40-370. While the N-terminal part of that minimal region (residues 47-247) folds into the prim/pol domain and bears the active site, the structure and function of the C-terminal part (residues 248-370) is unknown. Here we show that the C-terminal part of the minimal region folds into a compact domain with six helices and is stabilized by a disulfide bond. Three helices superimpose well with the C-terminal domain of the primase of the bacterial broad host range plasmid RSF1010. Structure-based site-directed mutagenesis shows that the C-terminal helix of the helix bundle domain is required for primase activity although it is distant to the active site in the crystallized conformation. Furthermore, we identified mutants of the C-terminal domain, which are defective in template binding, dinucleotide formation and conformation change prior to DNA extension.
- Institute of Biochemistry, University of Bayreuth, Universitätsstrasse 30, 95447 Bayreuth, Germany.
Organizational Affiliation: