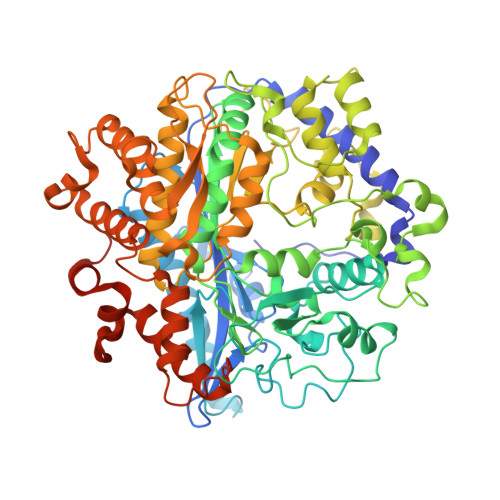
Structural basis for feedback and pharmacological inhibition of Saccharomyces cerevisiae glutamate cysteine ligase.
Biterova, E.I., Barycki, J.J.(2010) J Biological Chem 285: 14459-14466
- PubMed: 20220146 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M110.104802
- Primary Citation Related Structures:
3LVV, 3LVW - PubMed Abstract:
Structural characterization of glutamate cysteine ligase (GCL), the enzyme that catalyzes the initial, rate-limiting step in glutathione biosynthesis, has revealed many of the molecular details of substrate recognition. To further delineate the mechanistic details of this critical enzyme, we have determined the structures of two inhibited forms of Saccharomyces cerevisiae GCL (ScGCL), which shares significant sequence identity with the human enzyme. In vivo, GCL activity is feedback regulated by glutathione. Examination of the structure of ScGCL-glutathione complex (2.5 A; R = 19.9%, R(free) = 25.1%) indicates that the inhibitor occupies both the glutamate- and the presumed cysteine-binding site and disrupts the previously observed Mg(2+) coordination in the ATP-binding site. l-Buthionine-S-sulfoximine (BSO) is a mechanism-based inhibitor of GCL and has been used extensively to deplete glutathione in cell culture and in vivo model systems. Inspection of the ScGCL-BSO structure (2.2 A; R = 18.1%, R(free) = 23.9%) confirms that BSO is phosphorylated on the sulfoximine nitrogen to generate the inhibitory species and reveals contacts that likely contribute to transition state stabilization. Overall, these structures advance our understanding of the molecular regulation of this critical enzyme and provide additional details of the catalytic mechanism of the enzyme.
- Department of Biochemistry and the Redox Biology Center, University of Nebraska, Lincoln, Nebraska 68588, USA.
Organizational Affiliation: