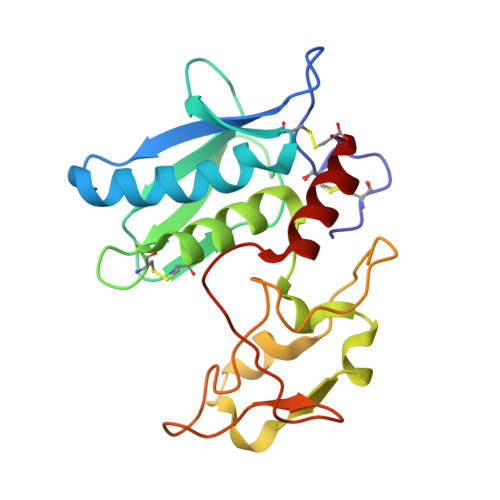
Crystal structure of zebrafish hatching enzyme 1 from the zebrafish Danio rerio
Okada, A., Sano, K., Nagata, K., Yasumasu, S., Ohtsuka, J., Yamamura, A., Kubota, K., Iuchi, I., Tanokura, M.(2010) J Mol Biology 402: 865-878
- PubMed: 20727360
- DOI: https://doi.org/10.1016/j.jmb.2010.08.023
- Primary Citation Related Structures:
3LQB - PubMed Abstract:
Fish hatching enzymes are zinc metalloproteases that digest the egg envelope (chorion) at the time of hatching. The crystal structure of zebrafish hatching enzyme 1 (ZHE1) has been solved at 1.10 Å resolution. ZHE1 is monomeric, is mitten shaped, and has a cleft at the center of the molecule. ZHE1 consists of three 3(10)-helices, three α-helices, and two β-sheets. The central cleft represents the active site of the enzyme that is crucial for substrate recognition and catalysis. Alanine-scanning mutagenesis of the two substrate peptides has shown that AspP1' contributes the most and that the residues at P4-P2' also contribute to the recognition of the major substrate peptide by ZHE1, whereas GluP3' and the hydrophobic residues at P4-P2, P2', and P5' contribute significantly to the recognition of the minor substrate peptide by ZHE1. Molecular models of these two substrate peptides bound to ZHE1 have been built based on the crystal structure of a transition-state analog inhibitor bound to astacin. In substrate-recognition models, the AspP1' in the major substrate peptide forms a salt bridge with Arg182 of ZHE1, while the GluP3' in the minor substrate peptide instead forms a salt bridge with Arg182. Thus, these two substrate peptides would be differently recognized by ZHE1. The shapes and electrostatic potentials of the substrate-binding clefts of ZHE1 and the structurally similar proteins astacin and bone morphogenetic protein 1 are significantly dissimilar due to different side chains, which would confer their distinctive substrate preferences.
- Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: