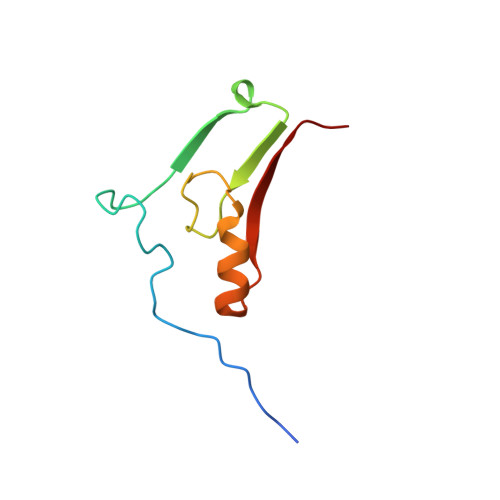
Mutation of the His ligand in mitoNEET stabilizes the 2Fe-2S cluster despite conformational heterogeneity in the ligand environment.
Conlan, A.R., Paddock, M.L., Homer, C., Axelrod, H.L., Cohen, A.E., Abresch, E.C., Zuris, J.A., Nechushtai, R., Jennings, P.A.(2011) Acta Crystallogr D Biol Crystallogr 67: 516-523
- PubMed: 21636891 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444911011577
- Primary Citation Related Structures:
3LPQ - PubMed Abstract:
MitoNEET is the only identified Fe-S protein localized to the outer mitochondrial membrane and a 1.5 Å resolution X-ray analysis has revealed a unique structure [Paddock et al. (2007), Proc. Natl Acad. Sci. USA, 104, 14342-14347]. The 2Fe-2S cluster is bound with a 3Cys-1His coordination which defines a new class of 2Fe-2S proteins. The hallmark feature of this class is the single noncysteine ligand His87, which when replaced by Cys decreases the redox potential (E(m)) by ∼300 mV and increases the stability of the cluster by around sixfold. Unexpectedly, the pH dependence of the lifetime of the 2Fe-2S cluster remains the same as in the wild-type protein. Here, the crystal structure of H87C mitoNEET was determined to 1.7 Å resolution (R factor = 18%) to investigate the structural basis of the changes in the properties of the 2Fe-2S cluster. In comparison to the wild type, structural changes are localized to the immediate vicinity of the cluster-binding region. Despite the increased stability, Cys87 displays two distinct conformations, with distances of 2.3 and 3.2 Å between the S(γ) and the outer Fe of the 2Fe-2S cluster. In addition, Lys55 exhibits multiple conformations in the H87C mutant protein. The structure and distinct characteristics of the H87C mutant provide a framework for further studies investigating the effects of mutation on the properties of the 2Fe-2S cluster in this new class of proteins.
- Departments of Chemistry and Biochemistry, University of California at San Diego, La Jolla, CA 92093, USA.
Organizational Affiliation: