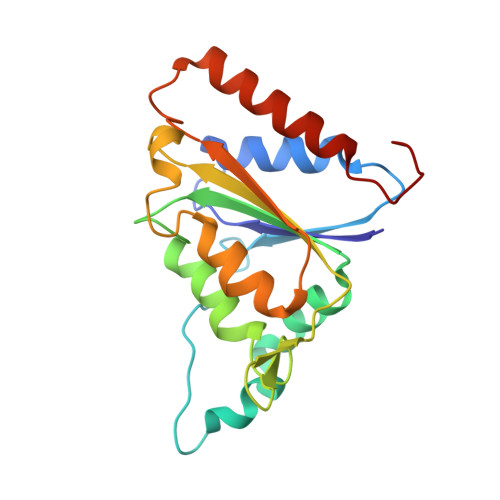
Structural and biochemical characterization of MdaB from cariogenic Streptococcus mutans reveals an NADPH-specific quinone oxidoreductase
Wang, Z.X., Li, L., Dong, Y.H., Su, X.-D.(2014) Acta Crystallogr D Biol Crystallogr 70: 912-921
- PubMed: 24699637 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004713033749
- Primary Citation Related Structures:
3LCM - PubMed Abstract:
The smu.1420 gene from the cariogenic pathogen Streptococcus mutans encodes a putative protein which has sequence homology to NQO [ quinone oxidoreductase] family members, including mammalian NQO and bacterial MdaB (modulator of drug activity B). NQO can detoxify quinones by converting them to hydroquinones and prevent the generation of reactive oxygen species. Thus, comprehensive studies on Smu.1420 will be important for uncovering the antioxidation and antidrug mechanisms of S. mutans. Here, the catalytic properties of Smu.1420 have been characterized, and its structure was determined in complexes with NADP(+) and menadione, respectively. Smu.1420 binds menadione directly and exhibits a pronounced preference for NADPH over NADH as a substrate, demonstrating that it is an NADPH-specific quinone oxidoreductase. The structure of Smu.1420 shows a compact homodimer with two substrate pockets located in the cleft of the dimer interface. The nicotinamide moiety of NADP(+) is bound on top of the isoalloxazine moiety of the FAD cofactor and overlaps with the binding site of menadione, suggesting a hydride-transfer process from NADPH to FAD and then to menadione. Two strongly basic patches near the substrate pocket are expected to confer the preference for NADPH over NADH. These studies shed light on future drug development against the cariogenic pathogen S. mutans.
- State Key Laboratory of Protein and Plant Gene Research, and Biodynamic Optical Imaging Center (BIOPIC), Peking University, Beijing 100871, People's Republic of China.
Organizational Affiliation: