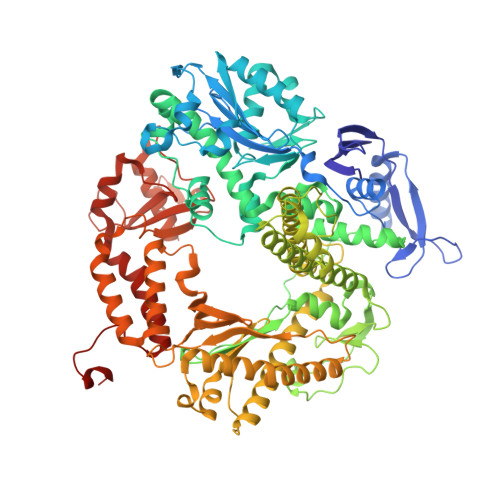
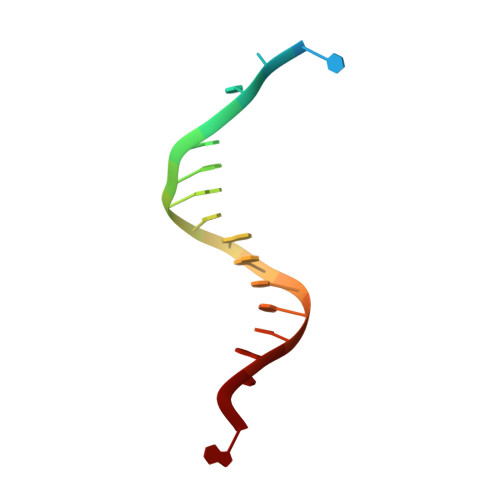
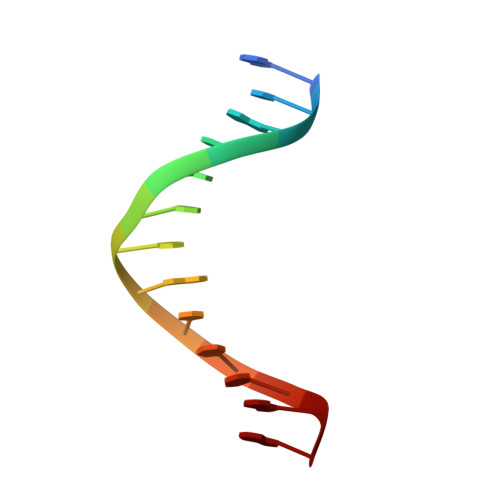
Crystal structure of a replicative DNA polymerase bound to the oxidized guanine lesion guanidinohydantoin.
Aller, P., Ye, Y., Wallace, S.S., Burrows, C.J., Doublie, S.(2010) Biochemistry 49: 2502-2509
- PubMed: 20166752 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi902195p
- Primary Citation Related Structures:
3L8B - PubMed Abstract:
The oxidation of guanine generates one of the most common DNA lesions, 8-oxo-7,8-dihydroguanine (8-oxoG). The further oxidation of 8-oxoG can produce either guanidinohydantoin (Gh) in duplex DNA or spiroiminodihydantoin (Sp) in nucleosides and ssDNA. Although Gh can be a strong block for replicative DNA polymerases such as RB69 DNA polymerase, this lesion is also mutagenic: DNA polymerases bypass Gh by preferentially incorporating a purine with a slight preference for adenine, which results in G.C --> T.A or G.C --> C.G transversions. The 2.15 A crystal structure of the replicative RB69 DNA polymerase in complex with DNA containing Gh reveals that Gh is extrahelical and rotated toward the major groove. In this conformation Gh is no longer in position to serve as a templating base for the incorporation of an incoming nucleotide. This work also constitutes the first crystallographic structure of Gh, which is stabilized in the R configuration in the two polymerase/DNA complexes present in the crystal asymmetric unit. In contrast to 8-oxoG, Gh is found in a high syn conformation in the DNA duplex and therefore presents the same hydrogen bond donor and acceptor pattern as thymine, which explains the propensity of DNA polymerases to incorporate a purine opposite Gh when bypass occurs.
- Department of Microbiology and Molecular Genetics, Stafford Hall, University of Vermont, Burlington, Vermont 05405, USA.
Organizational Affiliation: