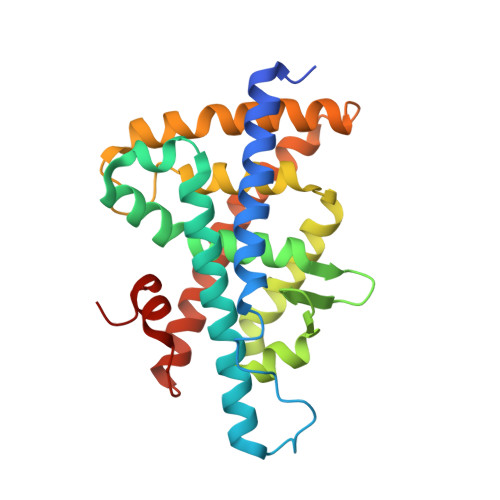
Discovery of tertiary sulfonamides as potent liver X receptor antagonists.
Zuercher, W.J., Buckholz, R.G., Campobasso, N., Collins, J.L., Galardi, C.M., Gampe, R.T., Hyatt, S.M., Merrihew, S.L., Moore, J.T., Oplinger, J.A., Reid, P.R., Spearing, P.K., Stanley, T.B., Stewart, E.L., Willson, T.M.(2010) J Med Chem 53: 3412-3416
- PubMed: 20345102 Search on PubMed
- DOI: https://doi.org/10.1021/jm901797p
- Primary Citation Related Structures:
3L0E - PubMed Abstract:
Tertiary sulfonamides were identified in a HTS as dual liver X receptor (LXR, NR1H2, and NR1H3) ligands, and the binding affinity of the series was increased through iterative analogue synthesis. A ligand-bound cocrystal structure was determined which elucidated key interactions for high binding affinity. Further characterization of the tertiary sulfonamide series led to the identification of high affinity LXR antagonists. GSK2033 (17) is the first potent cell-active LXR antagonist described to date. 17 may be a useful chemical probe to explore the cell biology of this orphan nuclear receptor.
- GlaxoSmithKline, Five Moore Drive, Research Triangle Park, North Carolina 27709, USA. william.j.zuercher@gsk.com
Organizational Affiliation: