Enzymatic and structural insights for substrate specificity of a family of jumonji histone lysine demethylases.
Horton, J.R., Upadhyay, A.K., Qi, H.H., Zhang, X., Shi, Y., Cheng, X.(2010) Nat Struct Mol Biol 17: 38-43
- PubMed: 20023638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1753
- Primary Citation Related Structures:
3KV4, 3KV5, 3KV6, 3KV9, 3KVA, 3KVB - PubMed Abstract:
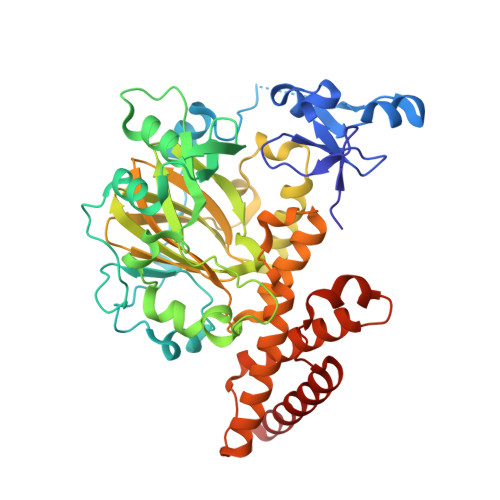
Combinatorial readout of multiple covalent histone modifications is poorly understood. We provide insights into how an activating histone mark, in combination with linked repressive marks, is differentially 'read' by two related human demethylases, PHF8 and KIAA1718 (also known as JHDM1D). Both enzymes harbor a plant homeodomain (PHD) that binds Lys4-trimethylated histone 3 (H3K4me3) and a jumonji domain that demethylates either H3K9me2 or H3K27me2. The presence of H3K4me3 on the same peptide as H3K9me2 makes the doubly methylated peptide a markedly better substrate of PHF8, whereas the presence of H3K4me3 has the opposite effect, diminishing the H3K9me2 demethylase activity of KIAA1718 without adversely affecting its H3K27me2 activity. The difference in substrate specificity between the two is explained by PHF8 adopting a bent conformation, allowing each of its domains to engage its respective target, whereas KIAA1718 adopts an extended conformation, which prevents its access to H3K9me2 by its jumonji domain when its PHD engages H3K4me3.
- Department of Biochemistry, Emory University School of Medicine, Atlanta, Georgia, USA.
Organizational Affiliation: