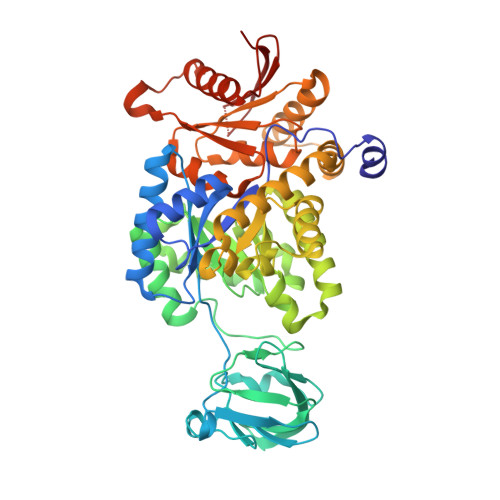
An improved strategy for the crystallization of Leishmania mexicana pyruvate kinase.
Morgan, H.P., McNae, I.W., Hsin, K.Y., Michels, P.A., Fothergill-Gilmore, L.A., Walkinshaw, M.D.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 215-218
- PubMed: 20208146 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109053494
- Primary Citation Related Structures:
3IS4, 3KTX - PubMed Abstract:
The inclusion of novel small molecules in crystallization experiments has provided very encouraging results and this method is now emerging as a promising alternative strategy for crystallizing 'problematic' biological macromolecules. These small molecules have the ability to promote lattice formation through stabilizing intermolecular interactions in protein crystals. Here, the use of 1,3,6,8-pyrenetetrasulfonic acid (PTS), which provides a helpful intermolecular bridge between Leishmania mexicana PYK (LmPYK) macromolecules in the crystal, is reported, resulting in the rapid formation of a more stable crystal lattice at neutral pH and greatly improved X-ray diffraction results. The refined structure of the LmPYK-PTS complex revealed the negatively charged PTS molecule to be stacked between positively charged (surface-exposed) arginine side chains from neighbouring LmPYK molecules in the crystal lattice.
- Structural Biochemistry Group, Institute of Structural and Molecular Biology, The University of Edinburgh, Michael Swann Building, The King's Buildings, Mayfield Road, Edinburgh EH9 3JR, Scotland.
Organizational Affiliation: