A human monoclonal antibody against insulin-like growth factor-II blocks the growth of human hepatocellular carcinoma cell lines in vitro and in vivo.
Dransfield, D.T., Cohen, E.H., Chang, Q., Sparrow, L.G., Bentley, J.D., Dolezal, O., Xiao, X., Peat, T.S., Newman, J., Pilling, P.A., Phan, T., Priebe, I., Brierley, G.V., Kastrapeli, N., Kopacz, K., Martik, D., Wassaf, D., Rank, D., Conley, G., Huang, Y., Adams, T.E., Cosgrove, L.(2010) Mol Cancer Ther 9: 1809-1819
- PubMed: 20515953 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-09-1134
- Primary Citation Related Structures:
3KR3 - PubMed Abstract:
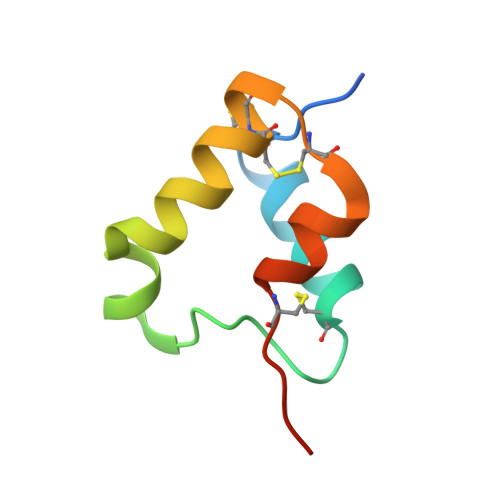
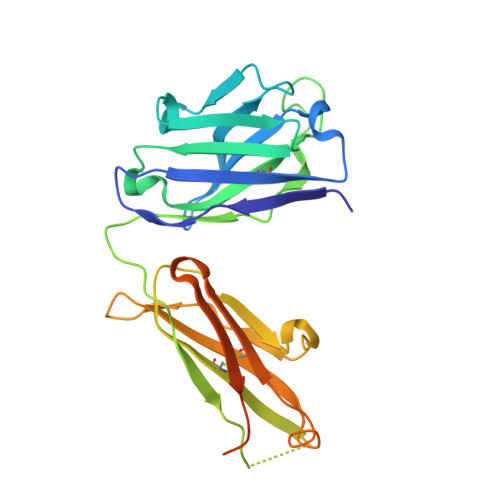
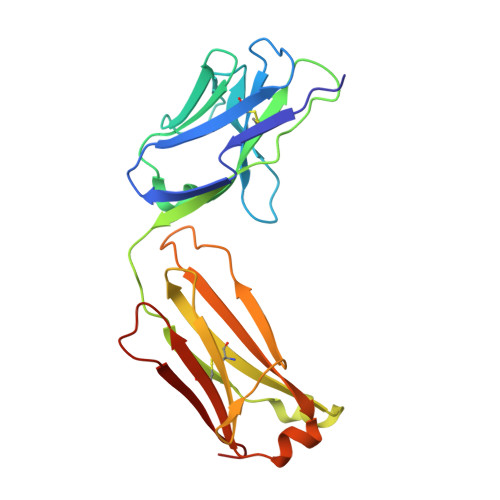
Elevated expression of insulin-like growth factor-II (IGF-II) is frequently observed in a variety of human malignancies, including breast, colon, and liver cancer. As IGF-II can deliver a mitogenic signal through both IGF-IR and an alternately spliced form of the insulin receptor (IR-A), neutralizing the biological activity of this growth factor directly is a potential alternative option to IGF-IR-directed agents. Using a Fab-displaying phage library and a biotinylated precursor form of IGF-II (1-104 amino acids) as a target, we isolated Fabs specific for the E-domain COOH-terminal extension form of IGF-II and for mature IGF-II. One of these Fabs that bound to both forms of IGF-II was reformatted into a full-length IgG, expressed, purified, and subjected to further analysis. This antibody (DX-2647) displayed a very high affinity for IGF-II/IGF-IIE (K(D) value of 49 and 10 pmol/L, respectively) compared with IGF-I (approximately 10 nmol/L) and blocked binding of IGF-II to IGF-IR, IR-A, a panel of insulin-like growth factor-binding proteins, and the mannose-6-phosphate receptor. A crystal complex of the parental Fab of DX-2647 bound to IGF-II was resolved to 2.2 A. DX-2647 inhibited IGF-II and, to a lesser extent, IGF-I-induced receptor tyrosine phosphorylation, cellular proliferation, and both anchorage-dependent and anchorage-independent colony formation in various cell lines. In addition, DX-2647 slowed tumor progression in the Hep3B xenograft model, causing decreased tumoral CD31 staining as well as reduced IGF-IIE and IGF-IR phosphorylation levels. Therefore, DX-2647 offers an alternative approach to targeting IGF-IR, blocking IGF-II signaling through both IGF-IR and IR-A.
- Discovery Research, Dyax Corp., Cambridge, Massachusetts 02139, USA. ddransfield@dyax.com
Organizational Affiliation: