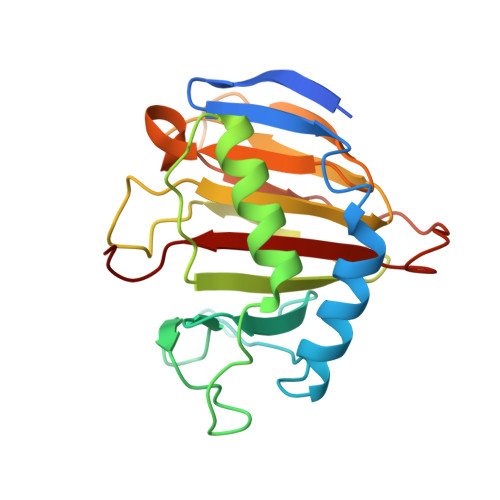
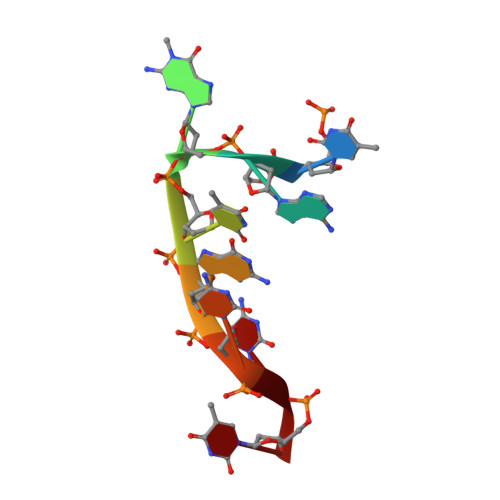
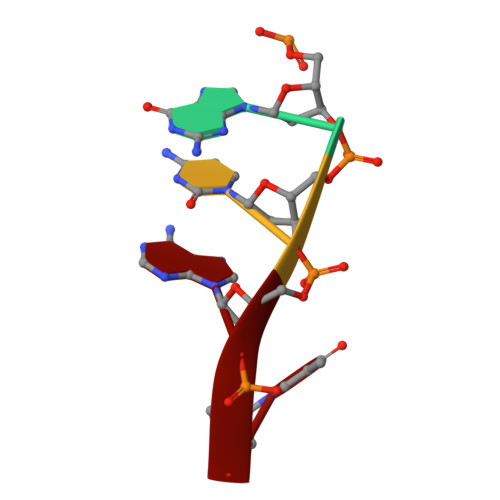
Structural and mutational analysis of Escherichia coli AlkB provides insight into substrate specificity and DNA damage searching.
Holland, P.J., Hollis, T.(2010) PLoS One 5: e8680-e8680
- PubMed: 20084272 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0008680
- Primary Citation Related Structures:
3KHB, 3KHC - PubMed Abstract:
In Escherichia coli, cytotoxic DNA methyl lesions on the N1 position of purines and N3 position of pyrimidines are primarily repaired by the 2-oxoglutarate (2-OG) iron(II) dependent dioxygenase, AlkB. AlkB repairs 1-methyladenine (1-meA) and 3-methylcytosine (3-meC) lesions, but it also repairs 1-methylguanine (1-meG) and 3-methylthymine (3-meT) at a much less efficient rate. How the AlkB enzyme is able to locate and identify methylated bases in ssDNA has remained an open question. We determined the crystal structures of the E. coli AlkB protein holoenzyme and the AlkB-ssDNA complex containing a 1-meG lesion. We coupled this to site-directed mutagenesis of amino acids in and around the active site, and tested the effects of these mutations on the ability of the protein to bind both damaged and undamaged DNA, as well as catalyze repair of a methylated substrate. A comparison of our substrate-bound AlkB-ssDNA complex with our unliganded holoenzyme reveals conformational changes of residues within the active site that are important for binding damaged bases. Site-directed mutagenesis of these residues reveals novel insight into their roles in DNA damage recognition and repair. Our data support a model that the AlkB protein utilizes at least two distinct conformations in searching and binding methylated bases within DNA: a "searching" mode and "repair" mode. Moreover, we are able to functionally separate these modes through mutagenesis of residues that affect one or the other binding state. Finally, our mutagenesis experiments show that amino acid D135 of AlkB participates in both substrate specificity and catalysis.
- Department of Biochemistry, Wake Forest University School of Medicine, Winston-Salem, North Carolina, USA.
Organizational Affiliation: