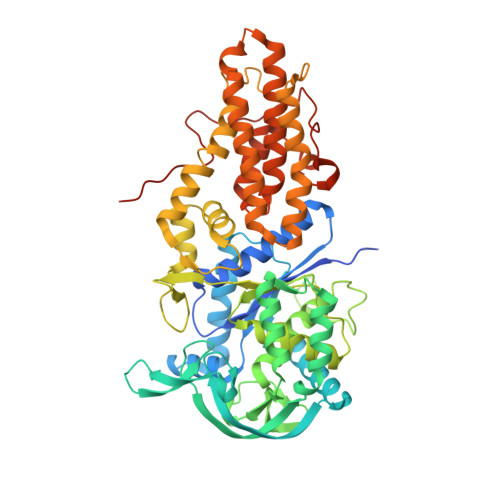
Structure of Leishmania major methionyl-tRNA synthetase in complex with intermediate products methionyladenylate and pyrophosphate.
Larson, E.T., Kim, J.E., Zucker, F.H., Kelley, A., Mueller, N., Napuli, A.J., Verlinde, C.L., Fan, E., Buckner, F.S., Van Voorhis, W.C., Merritt, E.A., Hol, W.G.(2011) Biochimie 93: 570-582
- PubMed: 21144880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.biochi.2010.11.015
- Primary Citation Related Structures:
3KFL - PubMed Abstract:
Leishmania parasites cause two million new cases of leishmaniasis each year with several hundreds of millions of people at risk. Due to the paucity and shortcomings of available drugs, we have undertaken the crystal structure determination of a key enzyme from Leishmania major in hopes of creating a platform for the rational design of new therapeutics. Crystals of the catalytic core of methionyl-tRNA synthetase from L. major (LmMetRS) were obtained with the substrates MgATP and methionine present in the crystallization medium. These crystals yielded the 2.0 Å resolution structure of LmMetRS in complex with two products, methionyladenylate and pyrophosphate, along with a Mg(2+) ion that bridges them. This is the first class I aminoacyl-tRNA synthetase (aaRS) structure with pyrophosphate bound. The residues of the class I aaRS signature sequence motifs, KISKS and HIGH, make numerous contacts with the pyrophosphate. Substantial differences between the LmMetRS structure and previously reported complexes of Escherichia coli MetRS (EcMetRS) with analogs of the methionyladenylate intermediate product are observed, even though one of these analogs only differs by one atom from the intermediate. The source of these structural differences is attributed to the presence of the product pyrophosphate in LmMetRS. Analysis of the LmMetRS structure in light of the Aquifex aeolicus MetRS-tRNA(Met) complex shows that major rearrangements of multiple structural elements of enzyme and/or tRNA are required to allow the CCA acceptor triplet to reach the methionyladenylate intermediate in the active site. Comparison with sequences of human cytosolic and mitochondrial MetRS reveals interesting differences near the ATP- and methionine-binding regions of LmMetRS, suggesting that it should be possible to obtain compounds that selectively inhibit the parasite enzyme.
- Department of Biochemistry, University of Washington, Seattle, WA 98195-7742, USA.
Organizational Affiliation: