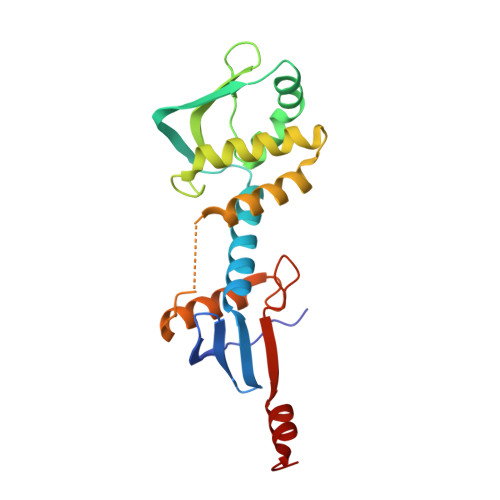
Structure of the endonuclease domain of MutL: unlicensed to cut.
Pillon, M.C., Lorenowicz, J.J., Uckelmann, M., Klocko, A.D., Mitchell, R.R., Chung, Y.S., Modrich, P., Walker, G.C., Simmons, L.A., Friedhoff, P., Guarne, A.(2010) Mol Cell 39: 145-151
- PubMed: 20603082
- DOI: https://doi.org/10.1016/j.molcel.2010.06.027
- Primary Citation of Related Structures:
3GAB, 3KDG, 3KDK - PubMed Abstract:
DNA mismatch repair corrects errors that have escaped polymerase proofreading, increasing replication fidelity 100- to 1000-fold in organisms ranging from bacteria to humans. The MutL protein plays a central role in mismatch repair by coordinating multiple protein-protein interactions that signal strand removal upon mismatch recognition by MutS. Here we report the crystal structure of the endonuclease domain of Bacillus subtilis MutL. The structure is organized in dimerization and regulatory subdomains connected by a helical lever spanning the conserved endonuclease motif. Additional conserved motifs cluster around the lever and define a Zn(2+)-binding site that is critical for MutL function in vivo. The structure unveils a powerful inhibitory mechanism to prevent undesired nicking of newly replicated DNA and allows us to propose a model describing how the interaction with MutS and the processivity clamp could license the endonuclease activity of MutL. The structure also provides a molecular framework to propose and test additional roles of MutL in mismatch repair.
- Department of Biochemistry and Biomedical Sciences, McMaster University, Hamilton, ON L8N 3Z5, Canada.
Organizational Affiliation: