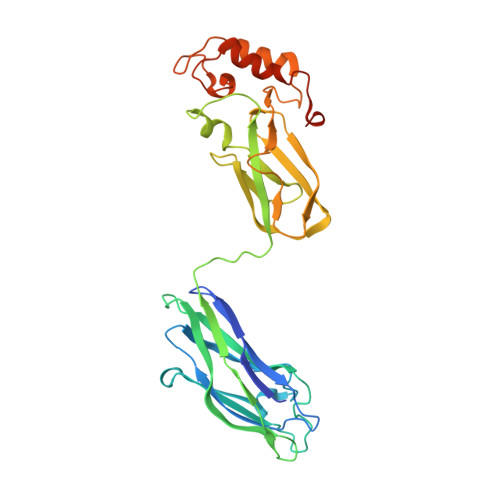
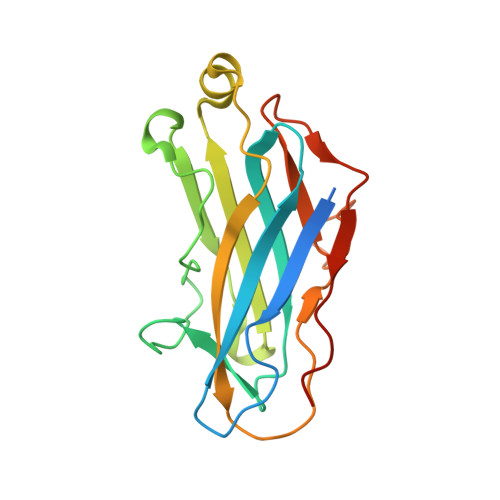
Insights into Higher-Order Organization of the Cellulosome Revealed by a Dissect-and-Build Approach: Crystal Structure of Interacting Clostridium thermocellum Multimodular Components
Adams, J.J., Currie, M.A., Ali, S., Bayer, E.A., Jia, Z., Smith, S.P.(2010) J Mol Biology 396: 833-839
- PubMed: 20070943 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2010.01.015
- Primary Citation Related Structures:
3KCP - PubMed Abstract:
Cellulosomes are large, multienzyme, plant cell wall-degrading protein complexes found affixed to the surface of a variety of anaerobic microbes. The core of the cellulosome is a noncatalytic scaffoldin protein, which contains several type-I cohesin modules that bind type-I dockerin-containing enzymatic subunits, a cellulose-binding module, an X module, and a type-II dockerin that interacts with type-II cohesin-containing cell surface proteins. The unique arrangement of the enzymatic subunits in the cellulosome complex, made possible by the scaffoldin subunit, promotes enhanced substrate degradation relative to the enzymes free in solution. Despite representative high-resolution structures of all of the individual modules of the cellulosome, this mechanism of enzymatic synergy remains poorly understood. Consequently, a model of the entire cellulosome and a detailed picture of intermodular contacts will provide more detailed insight into cellulosome activity. Toward this goal, we have solved the structure of a multimodular heterodimeric complex from Clostridium thermocellum composed of the type-II cohesin module of the cell surface protein SdbA bound to a trimodular C-terminal fragment of the scaffoldin subunit CipA to a resolution of 1.95 A. The linker that connects the ninth type-I cohesin module and the X module has elevated temperature factors, reflecting an inherent flexibility within this region. Interestingly, a novel dimer interface was observed between CipA and a second, symmetry-related CipA molecule within the crystal structure, mediated by contacts between a type-I cohesin and an X module of a symmetry mate, resulting in two intertwined scaffoldins. Sedimentation velocity experiments confirmed that dimerization also occurs in solution. These observations support the intriguing possibility that individual cellulosomes can associate with one another via inter-scaffoldin interactions, which may play a role in the mechanism of action of the complex.
- Department of Molecular and Cellular Physiology, Stanford University, Stanford, CA 94305, USA.
Organizational Affiliation: