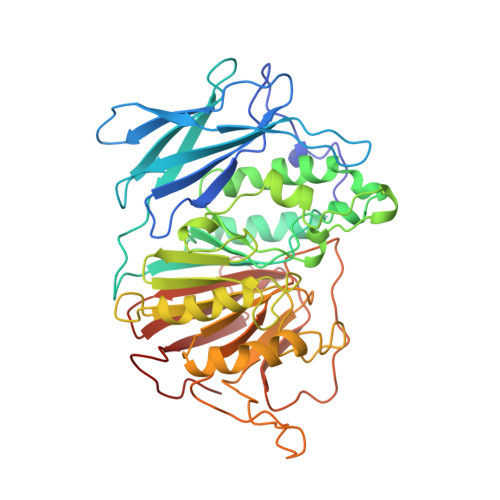
Mechanism of Fe(III)-Zn(II) purple acid phosphatase based on crystal structures.
Klabunde, T., Strater, N., Frohlich, R., Witzel, H., Krebs, B.(1996) J Mol Biology 259: 737-748
- PubMed: 8683579 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0354
- Primary Citation Related Structures:
1KBP, 3KBP, 4KBP - PubMed Abstract:
Purple acid phosphatase is a widely distributed non-specific phosphomonoesterase. X-ray structures of the dimeric 111-kDa Fe(III)-Zn(II) kidney bean purple acid phosphatase (kbPAP) complexed with phosphate, the product of the reaction, and with tungstate, a strong inhibitor of the phosphatase activity, were determined at 2.7 and 3.0 angstroms resolution, respectively. Furthermore the resolution of the unligated enzyme, recently solved at 2.9 angstroms could be extended to 2.65 angstroms with completely new data. The binding of both oxoanions is not accompanied by larger conformational changes in the enzyme structure. Small movements with a maximal coordinate shift of 1 angstroms are only observed for the active site residues His295 and His296. In the inhibitor complex as well as in the product complex, the oxoanion binds in a bidentate bridging mode to the two metal ions, replacing two of the presumed solvent ligands present in the unligated enzyme form. As also proposed for the unligated structure a bridging hydroxide ion completes the coordination spheres of both metal ions to octahedral arrangements. All three structures reported herein support a mechanism of phosphate ester hydrolysis involving interaction of the substrate with Zn(II) followed by a nucleophilic attack on the phosphorus by an Fe(III)-coordinated hydroxide ion. The negative charge evolving at the pentacoordinated transition state is probably stabilized by interactions with the divalent zinc and the imidazole groups of His202, His295, and His296, the latter protonating the leaving alcohol group.
- Anorganisch-Chemisches Institut, Universität Münster, Germany.
Organizational Affiliation: