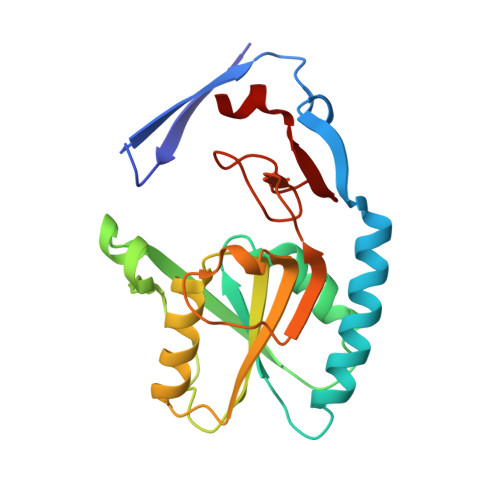
2.09 Angstrom resolution structure of a hypoxanthine-guanine phosphoribosyltransferase (hpt-1) from Bacillus anthracis str. 'Ames Ancestor' in complex with GMP
Halavaty, A.S., Minasov, G., Shuvalova, L., Dubrovska, I., Winsor, J., Papazisi, L., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.