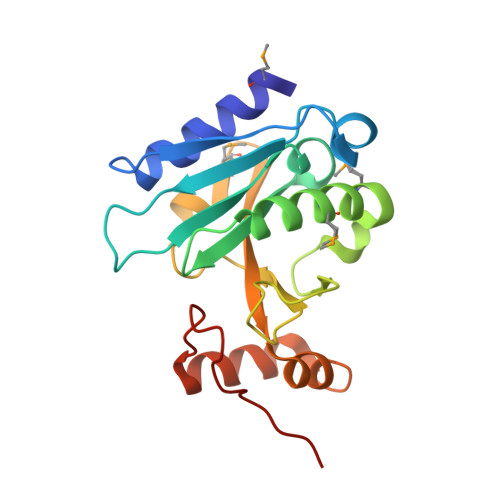
The crystal structure of a bacterial Sufu-like protein defines a novel group of bacterial proteins that are similar to the N-terminal domain of human Sufu.
Das, D., Finn, R.D., Abdubek, P., Astakhova, T., Axelrod, H.L., Bakolitsa, C., Cai, X., Carlton, D., Chen, C., Chiu, H.J., Chiu, M., Clayton, T., Deller, M.C., Duan, L., Ellrott, K., Farr, C.L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Kumar, A., Lam, W.W., Marciano, D., Miller, M.D., Morse, A.T., Nigoghossian, E., Nopakun, A., Okach, L., Puckett, C., Reyes, R., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Wooten, T., Xu, Q., Yeh, A., Zhou, J., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Protein Sci 19: 2131-2140
- PubMed: 20836087 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.497
- Primary Citation Related Structures:
3K5J - PubMed Abstract:
Sufu (Suppressor of Fused), a two-domain protein, plays a critical role in regulating Hedgehog signaling and is conserved from flies to humans. A few bacterial Sufu-like proteins have previously been identified based on sequence similarity to the N-terminal domain of eukaryotic Sufu proteins, but none have been structurally or biochemically characterized and their function in bacteria is unknown. We have determined the crystal structure of a more distantly related Sufu-like homolog, NGO1391 from Neisseria gonorrhoeae, at 1.4 Å resolution, which provides the first biophysical characterization of a bacterial Sufu-like protein. The structure revealed a striking similarity to the N-terminal domain of human Sufu (r.m.s.d. of 2.6 Å over 93% of the NGO1391 protein), despite an extremely low sequence identity of ∼15%. Subsequent sequence analysis revealed that NGO1391 defines a new subset of smaller, Sufu-like proteins that are present in ∼200 bacterial species and has resulted in expansion of the SUFU (PF05076) family in Pfam.
- Joint Center for Structural Genomics. http://www.jcsg.org
Organizational Affiliation: