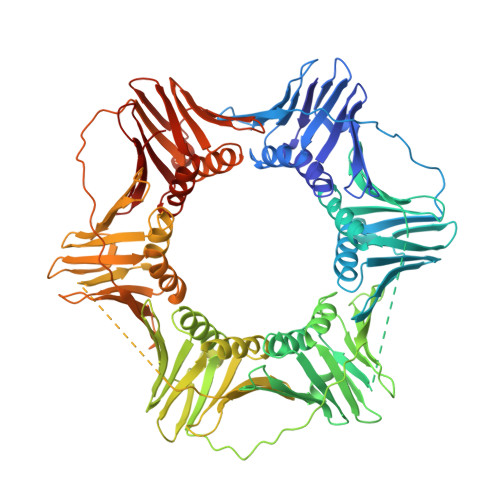
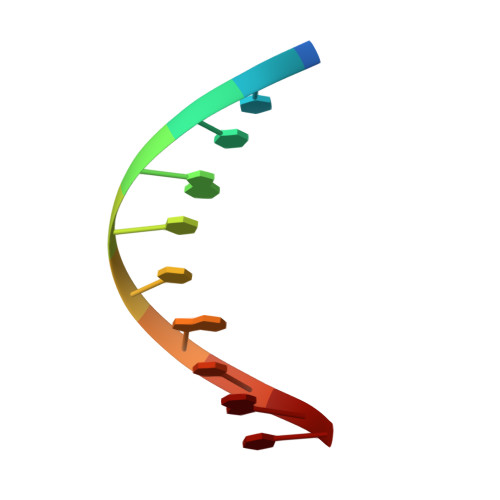
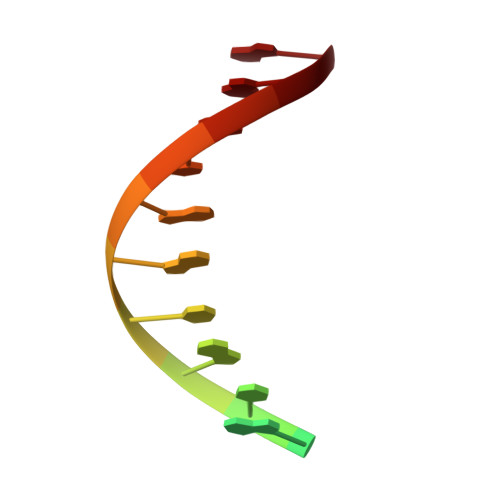
Analysis of the role of PCNA-DNA contacts during clamp loading.
McNally, R., Bowman, G.D., Goedken, E.R., O'Donnell, M., Kuriyan, J.(2010) BMC Struct Biol 10: 3-3
- PubMed: 20113510 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-10-3
- Primary Citation Related Structures:
3K4X - PubMed Abstract:
Sliding clamps, such as Proliferating Cell Nuclear Antigen (PCNA) in eukaryotes, are ring-shaped protein complexes that encircle DNA and enable highly processive DNA replication by serving as docking sites for DNA polymerases. In an ATP-dependent reaction, clamp loader complexes, such as the Replication Factor-C (RFC) complex in eukaryotes, open the clamp and load it around primer-template DNA. We built a model of RFC bound to PCNA and DNA based on existing crystal structures of clamp loaders. This model suggests that DNA would enter the clamp at an angle during clamp loading, thereby interacting with positively charged residues in the center of PCNA. We show that simultaneous mutation of Lys 20, Lys 77, Arg 80, and Arg 149, which interact with DNA in the RFC-PCNA-DNA model, compromises the ability of yeast PCNA to stimulate the DNA-dependent ATPase activity of RFC when the DNA is long enough to extend through the clamp. Fluorescence anisotropy binding experiments show that the inability of the mutant clamp proteins to stimulate RFC ATPase activity is likely caused by reduction in the affinity of the RFC-PCNA complex for DNA. We obtained several crystal forms of yeast PCNA-DNA complexes, measuring X-ray diffraction data to 3.0 A resolution for one such complex. The resulting electron density maps show that DNA is bound in a tilted orientation relative to PCNA, but makes different contacts than those implicated in clamp loading. Because of apparent partial disorder in the DNA, we restricted refinement of the DNA to a rigid body model. This result contrasts with previous analysis of a bacterial clamp bound to DNA, where the DNA was well resolved. Mutational analysis of PCNA suggests that positively charged residues in the center of the clamp create a binding surface that makes contact with DNA. Disruption of this positive surface, which had not previously been implicated in clamp loading function, reduces RFC ATPase activity in the presence of DNA, most likely by reducing the affinity of RFC and PCNA for DNA. The interaction of DNA is not, however, restricted to one orientation, as indicated by analysis of the PCNA-DNA co-crystals.
- Department of Molecular and Cell Biology, Department of Chemistry, California Institute for Quantitative Biosciences (QB3), Howard Hughes Medical Institute, University of California, Berkeley, Berkeley, CA 94720, USA.
Organizational Affiliation: