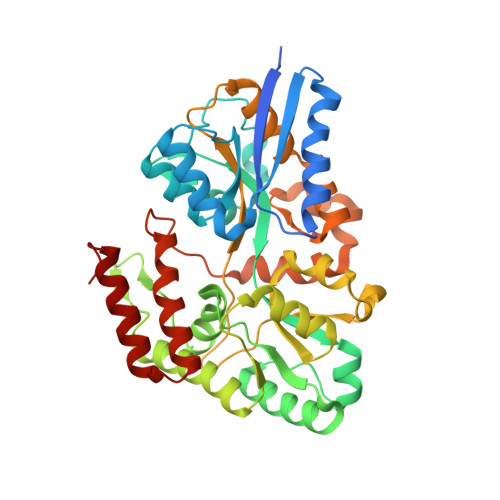
Crystal Structures of the Solute Receptor GacH of Streptomyces glaucescens in Complex with Acarbose and an Acarbose Homolog: Comparison with the Acarbose-Loaded Maltose-Binding Protein of Salmonella typhimurium.
Vahedi-Faridi, A., Licht, A., Bulut, H., Scheffel, F., Keller, S., Wehmeier, U.F., Saenger, W., Schneider, E.(2010) J Mol Biology 397: 709-723
- PubMed: 20132828
- DOI: https://doi.org/10.1016/j.jmb.2010.01.054
- Primary Citation of Related Structures:
3JYR, 3JZJ, 3K00, 3K01, 3K02 - PubMed Abstract:
GacH is the solute binding protein (receptor) of the putative oligosaccharide ATP-binding cassette transporter GacFG, encoded in the acarbose biosynthetic gene cluster (gac) from Streptomyces glaucescens GLA.O. In the context of the proposed function of acarbose (acarviosyl-1,4-maltose) as a 'carbophor,' the transporter, in complex with a yet to be identified ATPase subunit, is supposed to mediate the uptake of longer acarbose homologs and acarbose for recycling purposes. Binding assays using isothermal titration calorimetry identified GacH as a maltose/maltodextrin-binding protein with a low affinity for acarbose but with considerable binding activity for its homolog, component 5C (acarviosyl-1,4-maltose-1,4-glucose-1,1-glucose). In contrast, the maltose-binding protein of Salmonella typhimurium (MalE) displays high-affinity acarbose binding. We determined the crystal structures of GacH in complex with acarbose, component 5C, and maltotetraose, as well as in unliganded form. As found for other solute receptors, the polypeptide chain of GacH is folded into two distinct domains (lobes) connected by a hinge, with the interface between the lobes forming the substrate-binding pocket. GacH does not specifically bind the acarviosyl group, but displays specificity for binding of the maltose moiety in the inner part of its binding pocket. The crystal structure of acarbose-loaded MalE showed that two glucose units of acarbose are bound at the same region and position as maltose. A comparative analysis revealed that in GacH, acarbose is buried deeper into the binding pocket than in MalE by exactly one glucose ring shift, resulting in a total of 18 hydrogen-bond interactions versus 21 hydrogen-bond interactions for MalE(acarbose). Since the substrate specificity of ATP-binding cassette import systems is determined by the cognate binding protein, our results provide the first biochemical and structural evidence for the proposed role of GacHFG in acarbose metabolism.
- Institut für Chemie und Biochemie/Kristallographie, Freie Universität Berlin, Berlin, Germany.
Organizational Affiliation: