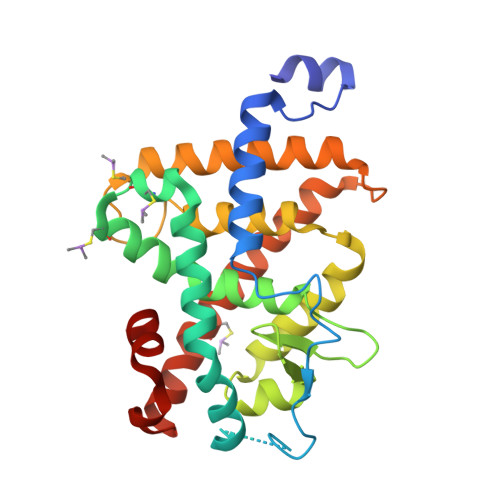
Gaining ligand selectivity in thyroid hormone receptors via entropy.
Martinez, L., Nascimento, A.S., Nunes, F.M., Phillips, K., Aparicio, R., Dias, S.M., Figueira, A.C., Lin, J.H., Nguyen, P., Apriletti, J.W., Neves, F.A., Baxter, J.D., Webb, P., Skaf, M.S., Polikarpov, I.(2009) Proc Natl Acad Sci U S A 106: 20717-20722
- PubMed: 19926848
- DOI: https://doi.org/10.1073/pnas.0911024106
- Primary Citation of Related Structures:
3JZB, 3JZC - PubMed Abstract:
Nuclear receptors are important targets for pharmaceuticals, but similarities between family members cause difficulties in obtaining highly selective compounds. Synthetic ligands that are selective for thyroid hormone (TH) receptor beta (TRbeta) vs. TRalpha reduce cholesterol and fat without effects on heart rate; thus, it is important to understand TRbeta-selective binding. Binding of 3 selective ligands (GC-1, KB141, and GC-24) is characterized at the atomic level; preferential binding depends on a nonconserved residue (Asn-331beta) in the TRbeta ligand-binding cavity (LBC), and GC-24 gains extra selectivity from insertion of a bulky side group into an extension of the LBC that only opens up with this ligand. Here we report that the natural TH 3,5,3'-triodothyroacetic acid (Triac) exhibits a previously unrecognized mechanism of TRbeta selectivity. TR x-ray structures reveal better fit of ligand with the TRalpha LBC. The TRbeta LBC, however, expands relative to TRalpha in the presence of Triac (549 A(3) vs. 461 A(3)), and molecular dynamics simulations reveal that water occupies the extra space. Increased solvation compensates for weaker interactions of ligand with TRbeta and permits greater flexibility of the Triac carboxylate group in TRbeta than in TRalpha. We propose that this effect results in lower entropic restraint and decreases free energy of interactions between Triac and TRbeta, explaining subtype-selective binding. Similar effects could potentially be exploited in nuclear receptor drug design.
- Instituto de Química, Universidade Estadual de Campinas, SP 13084-862, Campinas, Brazil.
Organizational Affiliation: