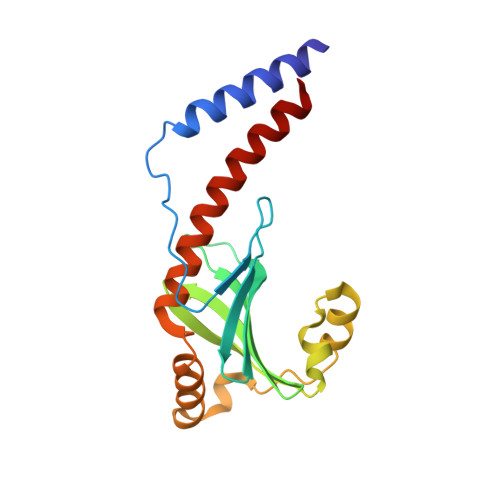
Structure of the Trypanosoma brucei p22 protein, a cytochrome oxidase subunit II-specific RNA-editing accessory factor.
Sprehe, M., Fisk, J.C., McEvoy, S.M., Read, L.K., Schumacher, M.A.(2010) J Biological Chem 285: 18899-18908
- PubMed: 20392699 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.066597
- Primary Citation Related Structures:
3JV1 - PubMed Abstract:
Kinetoplastid RNA (k-RNA) editing is a complex process in the mitochondria of kinetoplastid protozoa, including Trypanosoma brucei, that involves the guide RNA-directed insertion and deletion of uridines from precursor-mRNAs to produce mature, translatable mRNAs. k-RNA editing is performed by multiprotein complexes called editosomes. Additional non-editosome components termed k-RNA-editing accessory factors affect the extent of editing of specific RNAs or classes of RNAs. The T. brucei p22 protein was identified as one such accessory factor. Here we show that p22 contributes to cell growth in the procyclic form of T. brucei and functions as a cytochrome oxidase subunit II-specific k-RNA-editing accessory factor. To gain insight into its functions, we solved the crystal structure of the T. brucei p22 protein to 2.0-A resolution. The p22 structure consists of a six-stranded, antiparallel beta-sheet flanked by five alpha-helices. Three p22 subunits combine to form a tight trimer that is primarily stabilized by interactions between helical residues. One side of the trimer is strikingly acidic, while the opposite face is more neutral. Database searches show p22 is structurally similar to human p32, which has a number of functions, including regulation of RNA splicing. p32 interacts with a number of target proteins via its alpha1 N-terminal helix, which is among the most conserved regions between p22 and p32. Co-immunoprecipitation studies showed that p22 interacts with the editosome and the k-RNA accessory protein, TbRGG2, and alpha1 of p22 was shown to be important for the p22-TbRGG2 interaction. Thus, these combined studies suggest that p22 mediates its role in k-RNA editing by acting as an adaptor protein.
- Department of Biochemistry and Molecular Biology, University of Texas MD Anderson Cancer Center, Houston, Texas 77030, USA.
Organizational Affiliation: