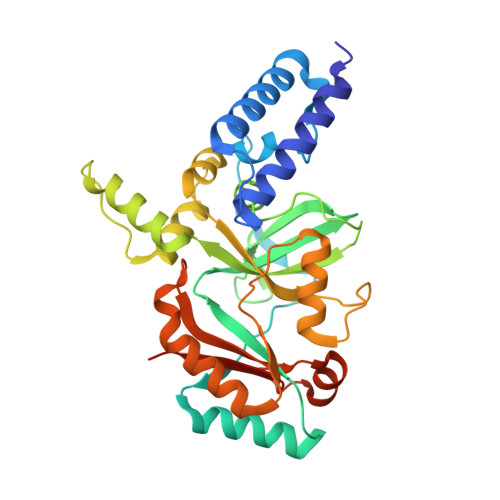
Structure of the adenylation domain of NAD(+)-dependent DNA ligase from Staphylococcus aureus.
Han, S., Chang, J.S., Griffor, M.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 1078-1082
- PubMed: 19923722 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109036872
- Primary Citation Related Structures:
3JSL, 3JSN - PubMed Abstract:
DNA ligase catalyzes phosphodiester-bond formation between immediately adjacent 5'-phosphate and 3'-hydroxyl groups in double-stranded DNA and plays a central role in many cellular and biochemical processes, including DNA replication, repair and recombination. Bacterial NAD(+)-dependent DNA ligases have been extensively characterized as potential antibacterial targets because of their essentiality and their structural distinction from human ATP-dependent DNA ligases. The high-resolution structure of the adenylation domain of Staphylococcus aureus NAD(+)-dependent DNA ligase establishes the conserved domain architecture with other bacterial adenylation domains. Two apo crystal structures revealed that the active site possesses the preformed NAD(+)-binding pocket and the 'C2 tunnel' lined with hydrophobic residues: Leu80, Phe224, Leu287, Phe295 and Trp302. The C2 tunnel is unique to bacterial DNA ligases and the Leu80 side chain at the mouth of the tunnel points inside the tunnel and forms a narrow funnel in the S. aureus DNA ligase structure. Taken together with other DNA ligase structures, the S. aureus DNA ligase structure provides a basis for a more integrated understanding of substrate recognition and catalysis and will be also be of help in the development of small-molecule inhibitors.
- Pfizer Inc., Groton, Connecticut 06340, USA. seungil.han@pfizer.com
Organizational Affiliation: