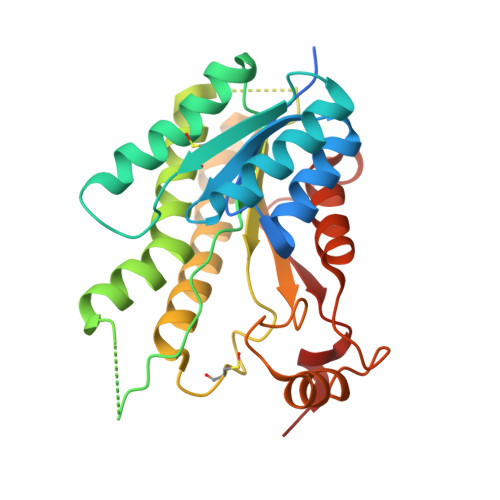
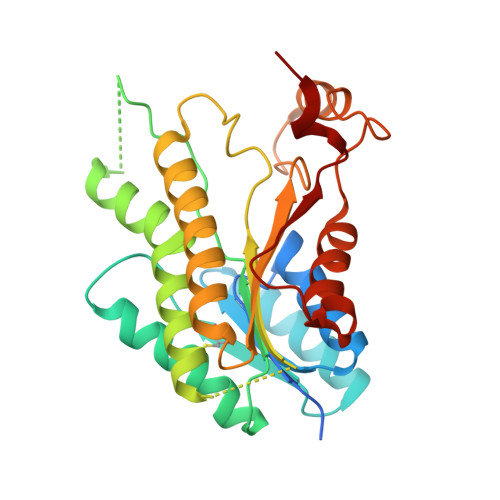
Structure-based design of pteridine reductase inhibitors targeting african sleeping sickness and the leishmaniases.
Tulloch, L.B., Martini, V.P., Iulek, J., Huggan, J.K., Lee, J.H., Gibson, C.L., Smith, T.K., Suckling, C.J., Hunter, W.N.(2010) J Med Chem 53: 221-229
- PubMed: 19916554 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm901059x
- Primary Citation Related Structures:
3BMC, 3BMN, 3BMO, 3BMQ, 3JQ6, 3JQ7, 3JQ8, 3JQ9, 3JQA, 3JQB, 3JQC, 3JQD, 3JQE, 3JQF, 3JQG - PubMed Abstract:
Pteridine reductase (PTR1) is a target for drug development against Trypanosoma and Leishmania species, parasites that cause serious tropical diseases and for which therapies are inadequate. We adopted a structure-based approach to the design of novel PTR1 inhibitors based on three molecular scaffolds. A series of compounds, most newly synthesized, were identified as inhibitors with PTR1-species specific properties explained by structural differences between the T. brucei and L. major enzymes. The most potent inhibitors target T. brucei PTR1, and two compounds displayed antiparasite activity against the bloodstream form of the parasite. PTR1 contributes to antifolate drug resistance by providing a molecular bypass of dihydrofolate reductase (DHFR) inhibition. Therefore, combining PTR1 and DHFR inhibitors might improve therapeutic efficacy. We tested two new compounds with known DHFR inhibitors. A synergistic effect was observed for one particular combination highlighting the potential of such an approach for treatment of African sleeping sickness.
- Division of Biological Chemistry and Drug Discovery, College of Life Sciences, University of Dundee, Dundee DD15EH, UK.
Organizational Affiliation: