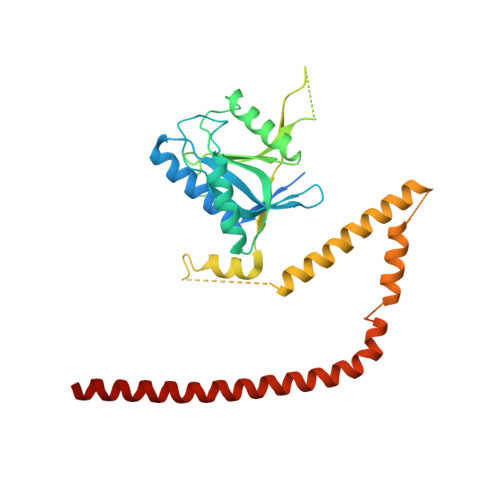
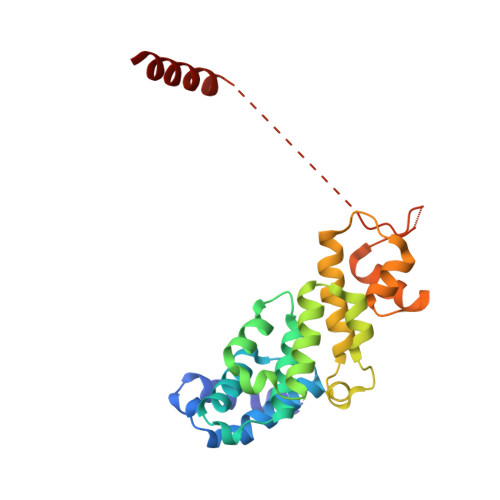
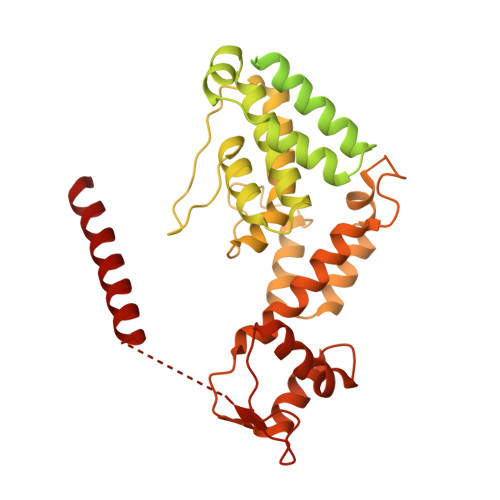
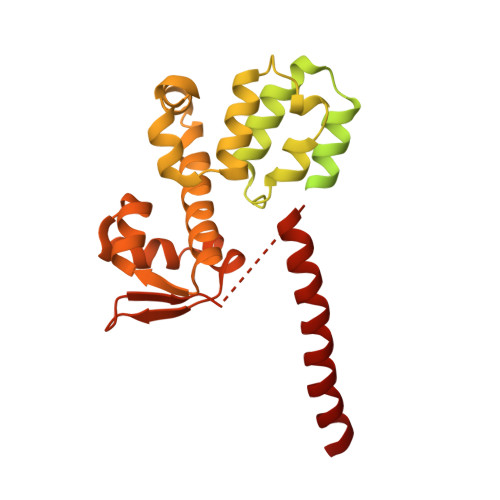
Molecular Architecture of the 40SeIF1eIF3 Translation Initiation Complex.
Erzberger, J.P., Stengel, F., Pellarin, R., Zhang, S., Schaefer, T., Aylett, C.H., Cimermancic, P., Boehringer, D., Sali, A., Aebersold, R., Ban, N.(2014) Cell 158: 1123-1135
- PubMed: 25171412 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2014.07.044
- Primary Citation Related Structures:
3J8B, 3J8C, 4U1C, 4U1D, 4U1E, 4U1F - PubMed Abstract:
Eukaryotic translation initiation requires the recruitment of the large, multiprotein eIF3 complex to the 40S ribosomal subunit. We present X-ray structures of all major components of the minimal, six-subunit Saccharomyces cerevisiae eIF3 core. These structures, together with electron microscopy reconstructions, cross-linking coupled to mass spectrometry, and integrative structure modeling, allowed us to position and orient all eIF3 components on the 40S⋅eIF1 complex, revealing an extended, modular arrangement of eIF3 subunits. Yeast eIF3 engages 40S in a clamp-like manner, fully encircling 40S to position key initiation factors on opposite ends of the mRNA channel, providing a platform for the recruitment, assembly, and regulation of the translation initiation machinery. The structures of eIF3 components reported here also have implications for understanding the architecture of the mammalian 43S preinitiation complex and the complex of eIF3, 40S, and the hepatitis C internal ribosomal entry site RNA.
- Department of Biology, Institute of Molecular Biology and Biophysics, ETH Zurich, Otto-Stern-Weg 5, 8093 Zurich, Switzerland. Electronic address: jan.erzberger@mol.biol.ethz.ch.
Organizational Affiliation: