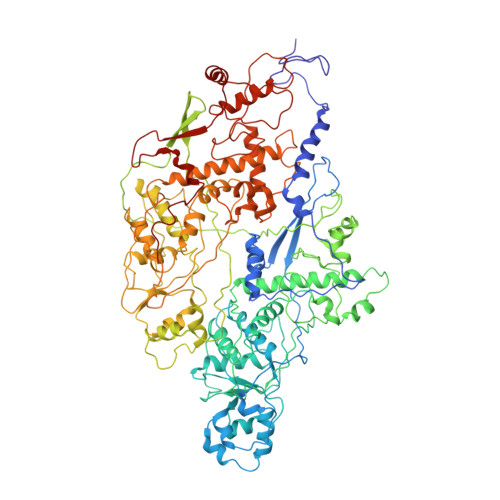
Cryo-EM near-atomic structure of a dsRNA fungal virus shows ancient structural motifs preserved in the dsRNA viral lineage.
Luque, D., Gomez-Blanco, J., Garriga, D., Brilot, A.F., Gonzalez, J.M., Havens, W.M., Carrascosa, J.L., Trus, B.L., Verdaguer, N., Ghabrial, S.A., Caston, J.R.(2014) Proc Natl Acad Sci U S A 111: 7641-7646
- PubMed: 24821769
- DOI: https://doi.org/10.1073/pnas.1404330111
- Primary Citation of Related Structures:
3J3I - PubMed Abstract:
Viruses evolve so rapidly that sequence-based comparison is not suitable for detecting relatedness among distant viruses. Structure-based comparisons suggest that evolution led to a small number of viral classes or lineages that can be grouped by capsid protein (CP) folds. Here, we report that the CP structure of the fungal dsRNA Penicillium chrysogenum virus (PcV) shows the progenitor fold of the dsRNA virus lineage and suggests a relationship between lineages. Cryo-EM structure at near-atomic resolution showed that the 982-aa PcV CP is formed by a repeated α-helical core, indicative of gene duplication despite lack of sequence similarity between the two halves. Superimposition of secondary structure elements identified a single "hotspot" at which variation is introduced by insertion of peptide segments. Structural comparison of PcV and other distantly related dsRNA viruses detected preferential insertion sites at which the complexity of the conserved α-helical core, made up of ancestral structural motifs that have acted as a skeleton, might have increased, leading to evolution of the highly varied current structures. Analyses of structural motifs only apparent after systematic structural comparisons indicated that the hallmark fold preserved in the dsRNA virus lineage shares a long (spinal) α-helix tangential to the capsid surface with the head-tailed phage and herpesvirus viral lineage.
- Department of Structure of Macromolecules, Centro Nacional de Biotecnología/Consejo Superior de Investigaciones Cientificas, Campus Cantoblanco, 28049 Madrid, Spain;Centro Nacional de Microbiología/Instituto de Salud Carlos III, 28220 Majadahonda, Madrid, Spain;
Organizational Affiliation: