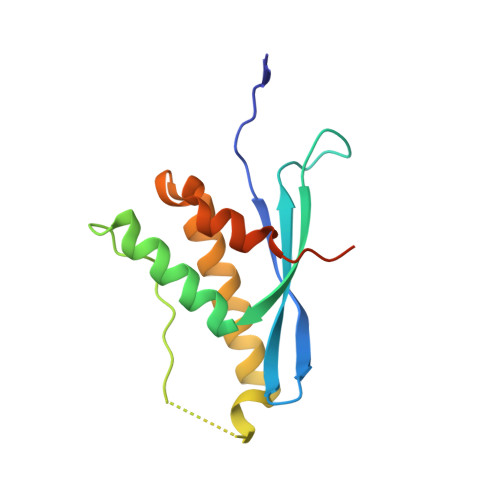
Human Sorting Nexin 7, Phox Homology (Px) Domain
Karlberg, T., Wisniewska, M., Arrowsmith, C.H., Berglund, H., Bountra, C., Collins, R., Edwards, A.M., Flodin, S., Flores, A., Graslund, S., Hammarstrom, M., Johansson, A., Johansson, I., Kallas, A., Kotenyova, T., Kotzsch, A., Kraulis, P., Moche, M., Nielsen, T.K., Nordlund, P., Nyman, T., Persson, C., Roos, A.K., Schutz, P., Siponen, M.I., Thorsell, A.G., Tresaugues, L., Van Den Berg, S., Weigelt, J., Welin, M., Schuler, H.To be published.