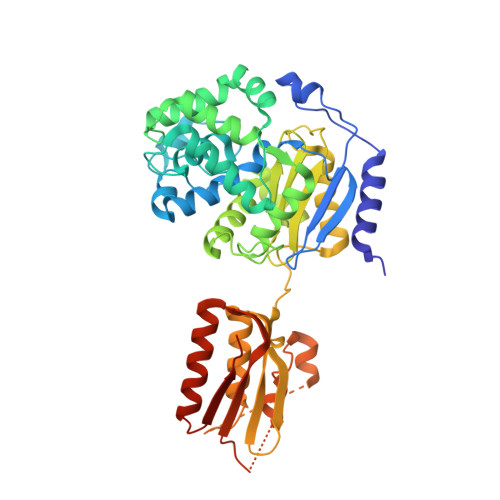
Crystal structure of salt-tolerant glutaminase from Micrococcus luteus K-3 in the presence and absence of its product l-glutamate and its activator Tris
Yoshimune, K., Shirakihara, Y., Wakayama, M., Yumoto, I.(2010) FEBS J 277: 738-748
- PubMed: 20050917 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2009.07523.x
- Primary Citation Related Structures:
3IF5, 3IH8, 3IH9, 3IHA, 3IHB - PubMed Abstract:
Glutaminase from Micrococcus luteus K-3 [Micrococcus glutaminase (Mglu); 456 amino acid residues (aa); 48 kDa] is a salt-tolerant enzyme. Our previous study determined the structure of its major 42-kDa fragment. Here, using new crystallization conditions, we determined the structures of the intact enzyme in the presence and absence of its product L-glutamate and its activator Tris, which activates the enzyme by sixfold. With the exception of a 'lid' part (26-29 aa) and a few other short stretches, the structures were all very similar over the entire polypeptide chain. However, the presence of the ligands significantly reduced the length of the disordered regions: 41 aa in the unliganded structure (N), 21 aa for L-glutamate (G), 8 aa for Tris (T) and 6 aa for both L-glutamate and Tris (TG). L-glutamate was identified in both the G and TG structures, whereas Tris was only identified in the TG structure. Comparison of the glutamate-binding site between Mglu and salt-labile glutaminase (YbgJ) from Bacillus subtilis showed significantly smaller structural changes of the protein part in Mglu. A comparison of the substrate-binding pocket of Mglu, which is highly specific for L-glutamine, with that of Erwinia carotovora asparaginase, which has substrates other than L-glutamine, shows that Mglu has a larger substrate-binding pocket that prevents the binding of L-asparagine with proper interactions.
- Research Institute of Genome-based Biofactory, National Institute of Advanced Industrial Science and Technology (AIST), Sapporo, Hokkaido, Japan. k.yoshimune@aist.go.jp
Organizational Affiliation: