A Catalytic Checkpoint in Base Excision by the Human 8-Oxoguanine DNA Glycosylase hOGG1
Crenshaw, C.M., Oo, K.S., Kutchukian, P.S., Verdine, G.L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| N-glycosylase/DNA lyase | 316 | Homo sapiens | Mutation(s): 1 Gene Names: MMH, MUTM, OGG1, OGH1 EC: 3.2.2 (PDB Primary Data), 4.2.99.18 (PDB Primary Data) | 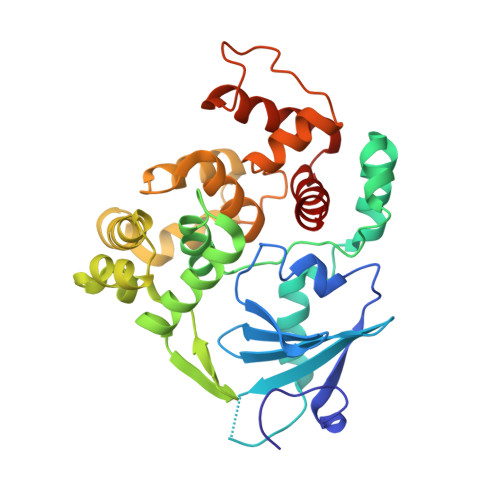 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: O15527 GTEx: ENSG00000114026 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O15527 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| 5'-D(*GP*GP*TP*AP*GP*AP*CP*CP*TP*GP*GP*AP*CP*G)-3' | 14 | N/A | 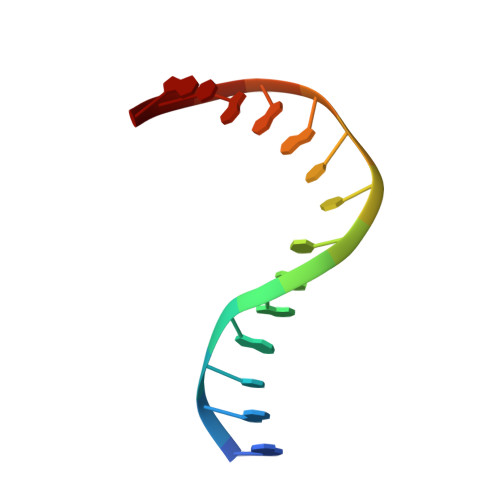 | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 91 | α = 90 |
| b = 91 | β = 90 |
| c = 211.6 | γ = 120 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| ADSC | data collection |
| DENZO | data reduction |
| SCALEPACK | data scaling |
| CNS | phasing |
| CNS | refinement |