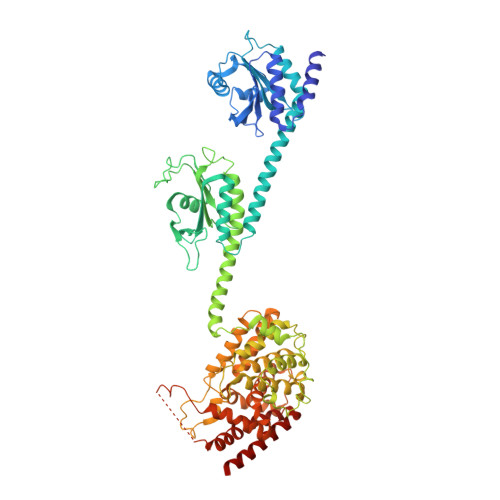
Mechanism for the allosteric regulation of phosphodiesterase 2A deduced from the X-ray structure of a near full-length construct.
Pandit, J., Forman, M.D., Fennell, K.F., Dillman, K.S., Menniti, F.S.(2009) Proc Natl Acad Sci U S A 106: 18225-18230
- PubMed: 19828435 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0907635106
- Primary Citation Related Structures:
3IBJ, 3ITM, 3ITU - PubMed Abstract:
We report the X-ray crystal structure of a phosphodiesterase (PDE) that includes both catalytic and regulatory domains. PDE2A (215-900) crystallized as a dimer in which each subunit had an extended organization of regulatory GAF-A and GAF-B and catalytic domains connected by long alpha-helices. The subunits cross at the GAF-B/catalytic domain linker, and each side of the dimer contains in series the GAF-A and GAF-B of one subunit and the catalytic domain of the other subunit. A dimer interface extends over the entire length of the molecule. The substrate binding pocket of each catalytic domain is occluded by the H-loop. We deduced from comparisons with structures of isolated, ligand-bound catalytic subunits that the H-loop swings out to allow substrate access. However, in dimeric PDE2A (215-900), the H-loops of the two catalytic subunits pack against each other at the dimer interface, necessitating movement of the catalytic subunits to allow for H-loop movement. Comparison of the unliganded GAF-B of PDE2A (215-900) with previous structures of isolated, cGMP-bound GAF domains indicates that cGMP binding induces a significant shift in the GAF-B/catalytic domain linker. We propose that cGMP binding to GAF-B causes movement, through this linker region, of the catalytic domains, such that the H-loops no longer pack at the dimer interface and are, instead, free to swing out to allow substrate access. This increase in substrate access is proposed as the basis for PDE2A activation by cGMP and may be a general mechanism for regulation of all PDEs.
- Pfizer Global Research and Development, Groton Labs, Groton, CT 06340, USA. jayvardhan.pandit@pfizer.com
Organizational Affiliation: