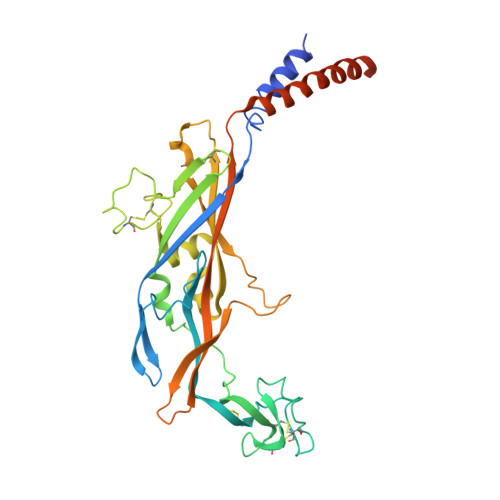
Crystal structure of the ATP-gated P2X(4) ion channel in the closed state.
Kawate, T., Michel, J.C., Birdsong, W.T., Gouaux, E.(2009) Nature 460: 592-598
- PubMed: 19641588 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature08198
- Primary Citation Related Structures:
3H9V, 3I5D - PubMed Abstract:
P2X receptors are cation-selective ion channels gated by extracellular ATP, and are implicated in diverse physiological processes, from synaptic transmission to inflammation to the sensing of taste and pain. Because P2X receptors are not related to other ion channel proteins of known structure, there is at present no molecular foundation for mechanisms of ligand-gating, allosteric modulation and ion permeation. Here we present crystal structures of the zebrafish P2X(4) receptor in its closed, resting state. The chalice-shaped, trimeric receptor is knit together by subunit-subunit contacts implicated in ion channel gating and receptor assembly. Extracellular domains, rich in beta-strands, have large acidic patches that may attract cations, through fenestrations, to vestibules near the ion channel. In the transmembrane pore, the 'gate' is defined by an approximately 8 A slab of protein. We define the location of three non-canonical, intersubunit ATP-binding sites, and suggest that ATP binding promotes subunit rearrangement and ion channel opening.
- Vollum Institute, Oregon Health and Science University, 3181 Southwest Sam Jackson Park Road, Oregon 97239, USA.
Organizational Affiliation: