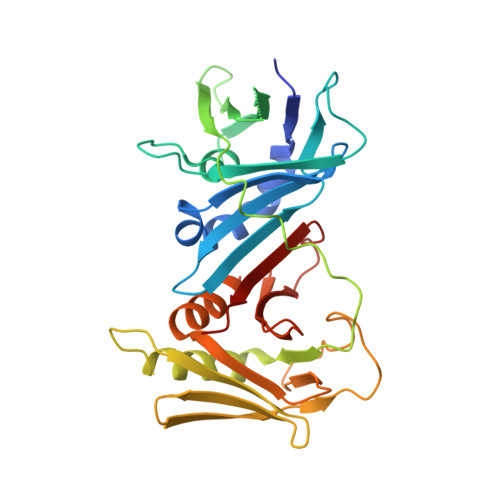
The crystal structure of PF-8, the DNA polymerase accessory subunit from Kaposi's sarcoma-associated herpesvirus.
Baltz, J.L., Filman, D.J., Ciustea, M., Silverman, J.E., Lautenschlager, C.L., Coen, D.M., Ricciardi, R.P., Hogle, J.M.(2009) J Virol 83: 12215-12228
- PubMed: 19759157 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01158-09
- Primary Citation Related Structures:
3HSL, 3I2M - PubMed Abstract:
Kaposi's sarcoma-associated herpesvirus is an emerging pathogen whose mechanism of replication is poorly understood. PF-8, the presumed processivity factor of Kaposi's sarcoma-associated herpesvirus DNA polymerase, acts in combination with the catalytic subunit, Pol-8, to synthesize viral DNA. We have solved the crystal structure of residues 1 to 304 of PF-8 at a resolution of 2.8 A. This structure reveals that each monomer of PF-8 shares a fold common to processivity factors. Like human cytomegalovirus UL44, PF-8 forms a head-to-head dimer in the form of a C clamp, with its concave face containing a number of basic residues that are predicted to be important for DNA binding. However, there are several differences with related proteins, especially in loops that extend from each monomer into the center of the C clamp and in the loops that connect the two subdomains of each protein, which may be important for determining PF-8's mode of binding to DNA and to Pol-8. Using the crystal structures of PF-8, the herpes simplex virus catalytic subunit, and RB69 bacteriophage DNA polymerase in complex with DNA and initial experiments testing the effects of inhibition of PF-8-stimulated DNA synthesis by peptides derived from Pol-8, we suggest a model for how PF-8 might form a ternary complex with Pol-8 and DNA. The structure and the model suggest interesting similarities and differences in how PF-8 functions relative to structurally similar proteins.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115, USA.
Organizational Affiliation: