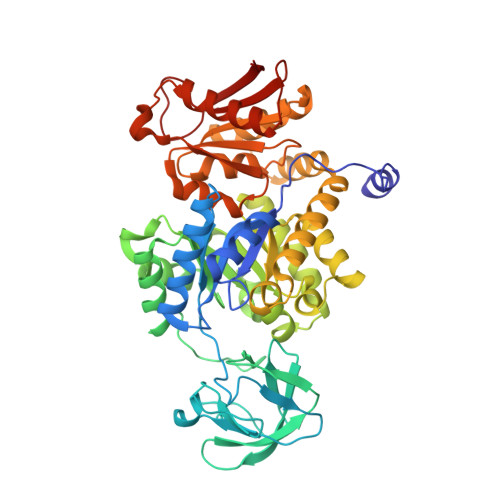
The allosteric mechanism of pryuvate kinase from Leishmania mexicana: a rock and lock model
Morgan, H.P., McNae, I.W., Nowicki, M.W., Hannaert, V., Michels, P.A.M., Fothergill-Gilmore, L.A., Walkinshaw, M.D.(2010) J Biological Chem 285: 12892-12898
- PubMed: 20123988
- DOI: https://doi.org/10.1074/jbc.M109.079905
- Primary Citation of Related Structures:
3HQN, 3HQO, 3HQP, 3HQQ - PubMed Abstract:
Allosteric regulation provides a rate management system for enzymes involved in many cellular processes. Ligand-controlled regulation is easily recognizable, but the underlying molecular mechanisms have remained elusive. We have obtained the first complete series of allosteric structures, in all possible ligated states, for the tetrameric enzyme, pyruvate kinase, from Leishmania mexicana. The transition between inactive T-state and active R-state is accompanied by a simple symmetrical 6 degrees rigid body rocking motion of the A- and C-domain cores in each of the four subunits. However, formation of the R-state in this way is only part of the mechanism; eight essential salt bridge locks that form across the C-C interface provide tetramer rigidity with a coupled 7-fold increase in rate. The results presented here illustrate how conformational changes coupled with effector binding correlate with loss of flexibility and increase in thermal stability providing a general mechanism for allosteric control.
- Structural Biochemistry Group, Institute of Structural and Molecular Biology, University of Edinburgh, Michael Swann Building, King's Buildings, Mayfield Road, Edinburgh EH9 3JR, Scotland, United Kingdom.
Organizational Affiliation: