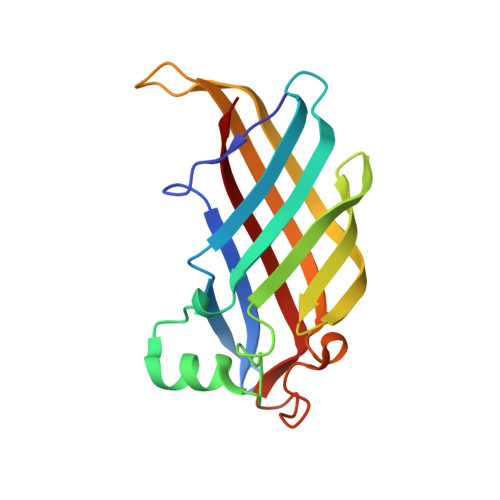
Helicobacter pylori acidic stress response factor HP1286 is a YceI homolog with new binding specificity.
Sisinni, L., Cendron, L., Favaro, G., Zanotti, G.(2010) FEBS J 277: 1896-1905
- PubMed: 20236316 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2010.07612.x
- Primary Citation Related Structures:
3HPE - PubMed Abstract:
HP1286 from Helicobacter pylori is among the proteins that play a relevant role in bacterial colonization and persistence in the stomach. Indeed, it was demonstrated to be overexpressed under acidic stress conditions, together with other essential virulence factors. Here we describe its crystal structure, determined at 2.1 A resolution. The molecular model, a dimer characterized by two-fold symmetry, shows that HP1286 structurally belongs to the YceI-like protein family, which in turn is characterized by the lipocalin fold. The latter characterizes proteins possessing an internal cavity with the function of binding and/or transport of amphiphilic molecules. Surprisingly, a molecule of erucamide was found bound in the internal cavity of each monomer of recombinant HP1286, cloned and expressed in an Escherichia coli heterologous system. The shape and length of the cavity indicate that, at variance with other members of the family, HP-YceI has a binding specificity for amphiphilic compounds with a linear chain of about 22 carbon atoms. These features, along with the fact that the protein is secreted by the bacterium and is involved in adaptation to an acidic environment, suggest that its function could be that of sequestering specific fatty acids or amides from the environment, either to supply the bacterium with the fatty acids necessary for its metabolism, or to protect and detoxify it from the detergent-like antimicrobial activity of fatty acids that are eventually present in the external milieu.
- Department of Biological Chemistry, University of Padua, Padua, Italy.
Organizational Affiliation: